Introduction
During the progression of advanced tumors, the body is often in a hypercoagulable state for various reasons. Cancer-related thrombosis is the leading cause of non-cancer mortality in malignancies [1]. In addition, coagulation-related factors also seem to influence the occurrence and development of tumors and the remodeling of microenvironment. Studies have confirmed that platelets can promote the distant metastasis of tumor cells by promoting epithelial-mesenchymal transformation of tumor cells [2]. In a research of colorectal cancer, the researchers found that an increase in fibrinogen levels was positively correlated with colorectal cancer as an independent risk factor [3]. Thrombin-related factors are closely related to tumor angiogenesis and tumor migration [4]. However, the current evidence mainly focuses on basic experiments and animal experiments, and has not been studied in clinical cohorts. In addition, the important role of coagulation-related factors in bladder cancer has not been reported yet.
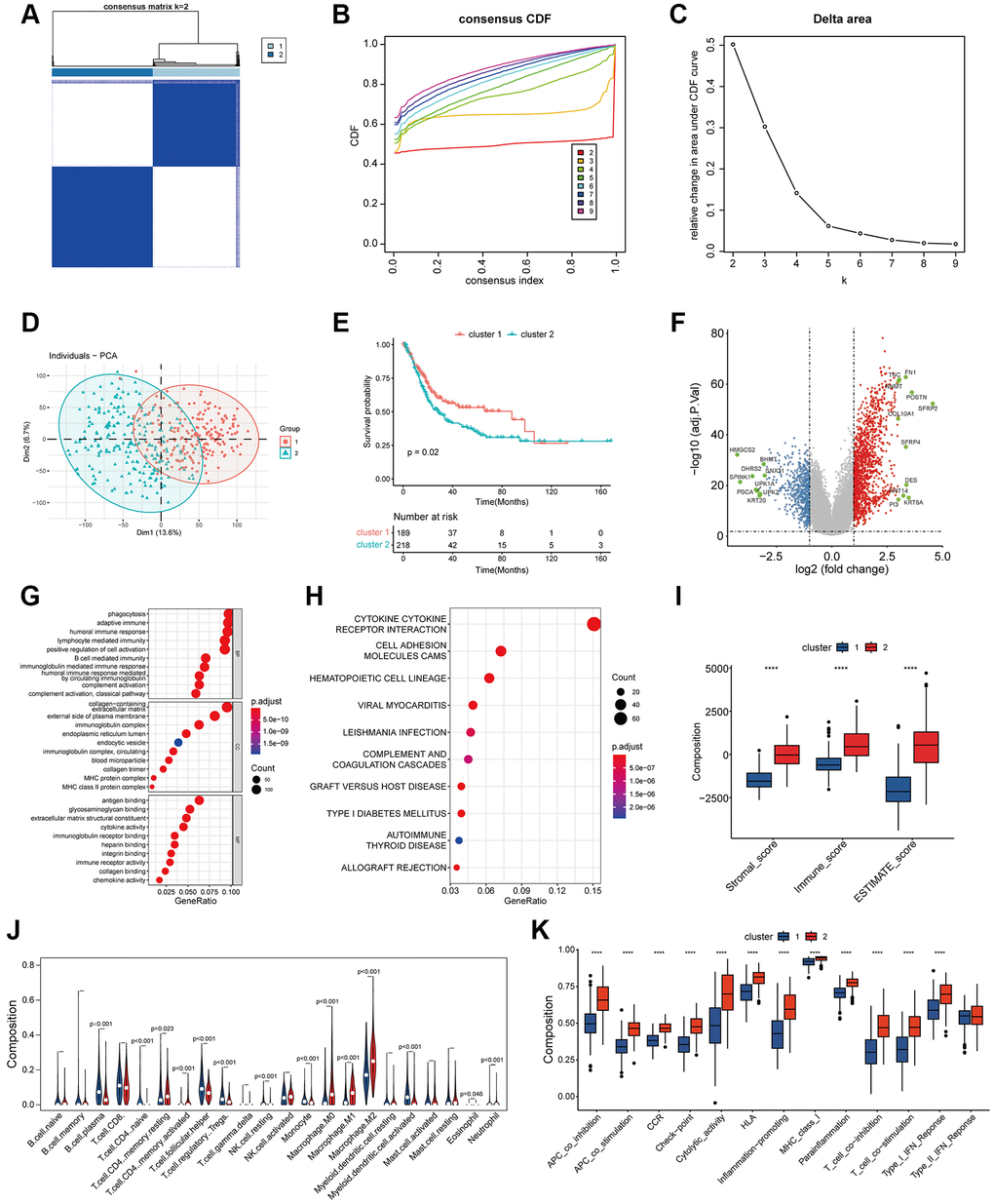
It has been proved feasible to construct a risk signature with the expression of key genes to monitor the tumor and its microenvironment state [5–7]. We divided bladder cancer into two states of hypercoagulability and low coagulation by clustering coagulation-related genes, and we found that hypercoagulability presaged a poor prognosis of bladder tumors. Through further screening for coagulation-related differentially expressed genes and constructing risk signature, we found that hypercoagulable state is closely related to tumor angiogenesis, cancer cell proliferation, EMT transformation, and immune microenvironment remodeling.
Materials and Methods
Patients and cohorts
The bladder cancer cohort from the Cancer Genome Atlas (TCGA) and GSE13057 database included information from 406 and 165 bladder cancer patients, respectively [8, 9]. The bladder cancer data set treated with PD1/PDL1 was derived from the Imvigor 210 data set and the GSE176307 data set for 298 and 88 patients, respectively [10, 11]. TCGA, GEO and Imvigor 210 are public databases in which patients have been ethically approved. Our research is based on open source data, so there are no ethical issues or other conflicts of interest. In our data screening process, cases with clinical information including survival state, survival time, clinical grade and stage, and complete information on genomic RNA sequence data met the inclusion criteria. Patients with incomplete information were removed. Data of the patients were downloaded through the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/) and TCGA database (https://portal.gdc.cancer.gov/repository). The data of Imvigor 210 was down were downloaded from (http://research-pub.gene.com/IMvigor210CoreBiologies). All genomic RNA sequence data were converted to transcripts per kilobase of exon model per million mapped reads (TPM), and standardize it by log2 [12].
Analysis of lymphocyte infiltration in tumor immune microenvironment
We calculated the infiltrating abundance of lymphocytes in each tumor sample using the cibersort deconvolution algorithm (CIBERSORT (https://cibersort.stanford.edu/) and its own LM22 lymphocyte genetic characteristics [20]. The obtained results are filtered (p < 0.05). The difference of infiltration between subgroups was evaluated by rank sum test, and the correlation between infiltration degree and risk score was evaluated by Pearson correlation.
Functional and pathway enrichment analysis
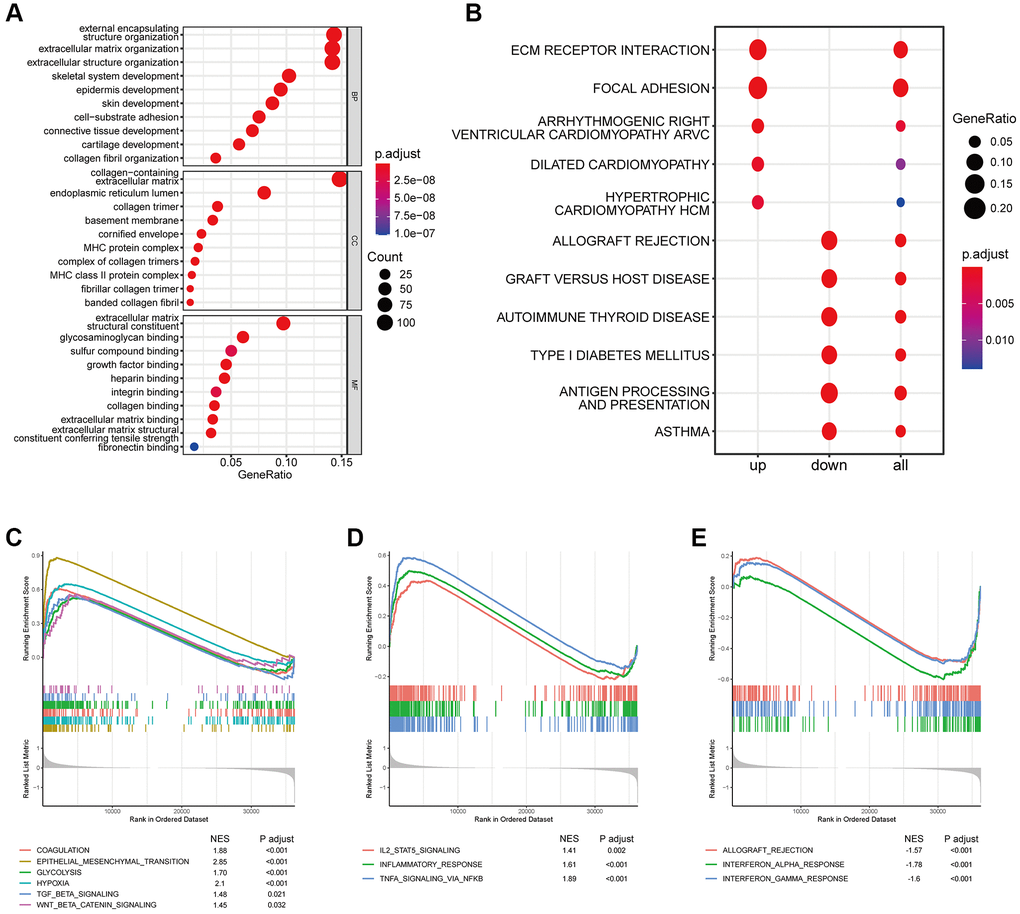
Coagulation-related signature was analyzed using the Cluster Profiler software package for gene annotation enrichment analysis [21]. The Kyoto Encyclopedia of Genes and Genomes (KEGG) and the Gene Ontology (GO) functional enrichment analyses were carried out to clarify the biological function and behavior of coagulation-related signature. The GSEA was used to show the potential pathways and mechanisms between high-risk and low-risk subgroups.
Statistical analysis
R software (R version 4.0.2 (2020-06-22)) was used for all analyses and graphics generated in this study. Inter-group comparisons were performed using the Kruskal-Wallis and t tests. If the normal distribution is satisfied, t-test is used. If the distribution is non-normal, Kruskal-Wallis test is used. Correlations were analyzed using the and Spearman correlation coefficient. Log-rank test, Cox regression and Kaplan-Meier curve were used to evaluate the prognostic value. All statistical analyses were bilateral, P-values < 0.05 were considered statistically significant.
Results
Establishment and verification of coagulation risk signature
We simultaneously constructed a coagulation-related risk signature by screening differentially expressed genes in different coagulation subtypes of bladder cancer and using the lasso-cox method (Figure 2A–2C). We split the TCGA data set into a training set and an internal verification set, and preliminarily verified the obtained risk signature. Our risk signature provides good predictability in both data sets. The area under the curve (AUC) in the training set was 0.76, and the KM curve suggested that the prognosis of patients in the high-risk and low-risk groups was significantly distinguished (HR = 3.28, p < 0.001). The AUC of the internal validation set was 0.74, and the prognosis of patients in the high-risk and low-risk groups was also well distinguished (HR = 3.5, p < 0.001). (Figure 2D, 2E). In order to further verify the sensitivity of our classifier, we made an in-depth analysis of the data of 165 bladder cancer patients in GSE13507 as an external validation set. In the overall data set of TCGA, the prognosis of patients in high and low risk groups is well differentiated (HR = 3.43, p < 0.001), and the AUC of one year, three years and five years are 0.75, 0.79 and 0.75 respectively (Figure 3A–3C). In the external validation set, the difference of prognosis between high and low risk groups was also statistically significant (HR = 1.67, p = 0.031), and the AUC of one year, three years and five years were 0.77, 0.64 and 0.6 respectively (Figure 3D–3F). Multivariate regression results suggested that our risk signature was an independent prognostic factor, whether in the TCGA dataset or in the external validation set (Figure 3G, 3H).
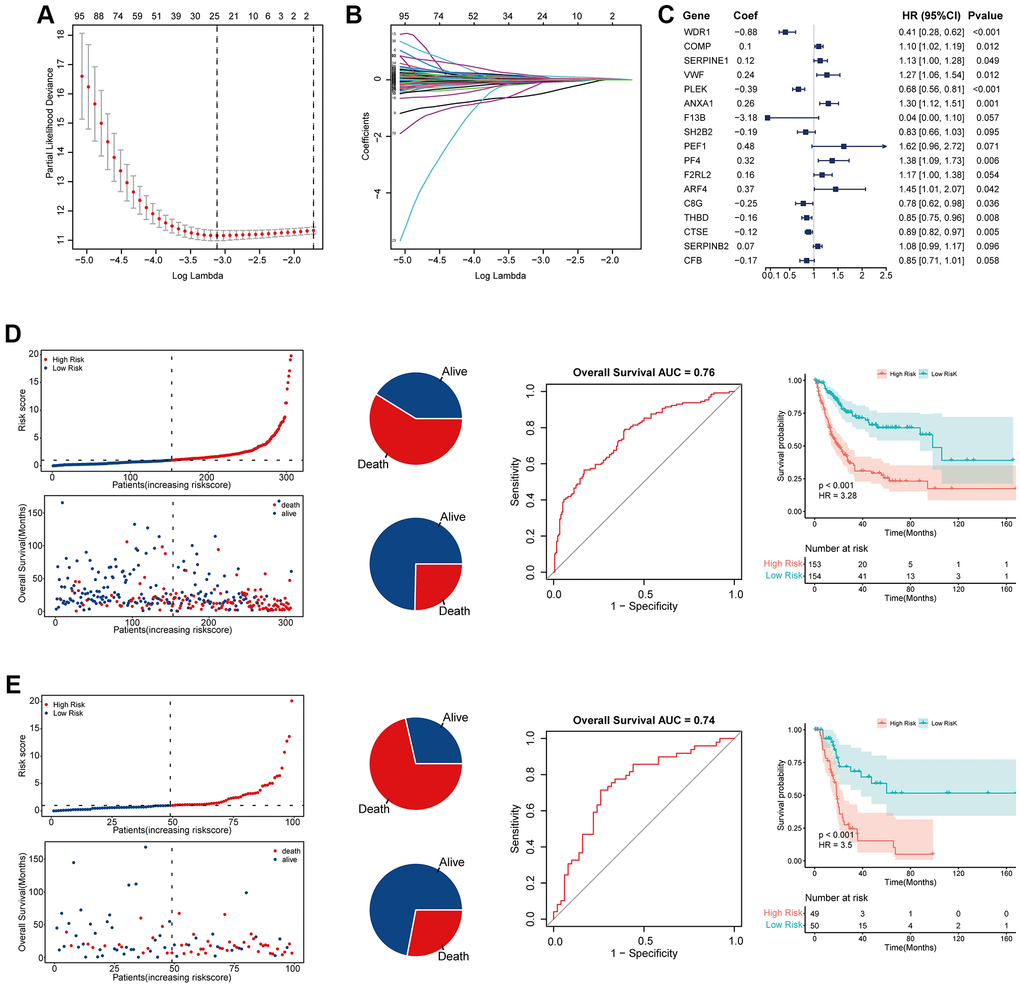
Figure 2. Establishment and validation of coagulation-related risk signature. (A) Least absolute shrinkage and selection operator cox analysis of the DEGs. (B) Partial likelihood deviance coefficient profiles. (C) Forest map of the variables contained in the risk signature. (D) Risk factor diagram, ROC curve and KM curve of the risk signature in the training set. The AUC of ROC curve is 0.76, while HR of KM curve is 3.28, p < 0.001. (E) Risk factor diagram, ROC curve and KM curve of the risk signature in the internal test set. The AUC of ROC curve is 0.74, while HR of KM curve is 3.5, p < 0.001.
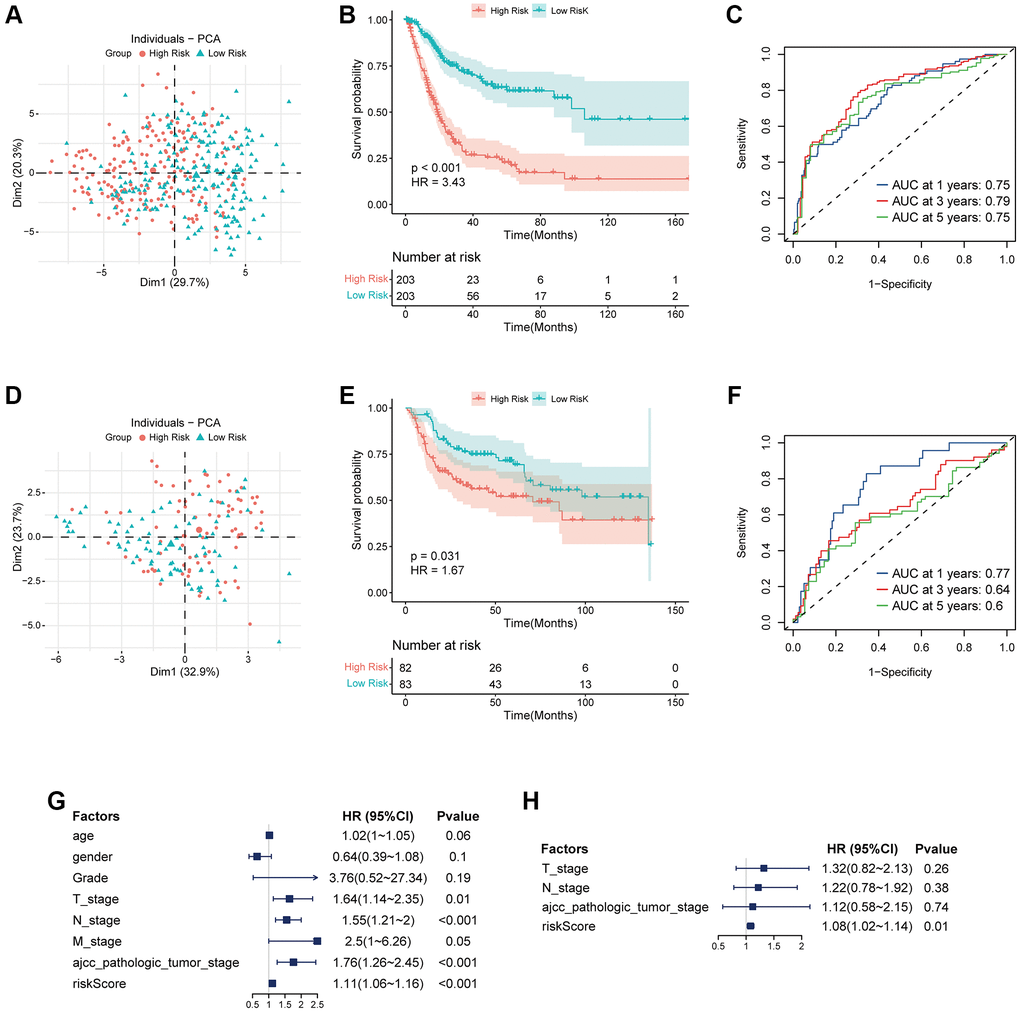
Figure 3. Validation of coagulation-related risk signature in TCGA set and external validation set. (A–C) PCA classification map, K-M curve, and ROC curve in TCGA dataset. The 1-year, 3-year, 5-year AUC of ROC curve is 0.75, 0.79, 0.75, respectively, while HR of KM curve is 3.43, p < 0.001. (D–F) PCA classification map, K-M curve, and ROC curve in external validation dataset. The 1-year, 3-year, 5-year AUC of ROC curve is 0.77, 0.64, 0.6, respectively, while HR of KM curve is 1.67, p = 0.031. (G) Multivariate COX regression forest map in TCGA dataset. Riskscore is an independent risk factor for prognosis. HR = 1.11, p < 0.001. (H) Multivariate COX regression forest map in external validation dataset. Riskscore is an independent risk factor for prognosis. HR = 1.08, p = 0.01.
Discussion
Coagulation abnormalities are very common in advanced tumors, and thromboembolism is the second leading cause of death in patients with advanced tumors, with a incidence rate only lower than organ failure due to distant metastasis of the tumor [23]. About 20% of symptomatic patients with deep vein thrombosis have active malignancies, and the incidence of thrombosis in neoplastic patients is 4–7 times higher than in non-neoplastic patients [24, 25]. Hypercoagulable state in patients with advanced tumor is formed by a combination of many factors. In addition to the factors related to the patients themselves, such as older age and obesity, tumor-related factors are also one of the causes. Studies have confirmed that tumors can secrete inflammatory factors and tumor procoagulant factors activate the clotting mechanism of the body. The cancer cell procoagulant directly promotes the conversion of factor X to factor Xa [26]. The sialic acid portion of the adenocarcinoma mucin causes direct non-enzymatic activation of factor X [27].
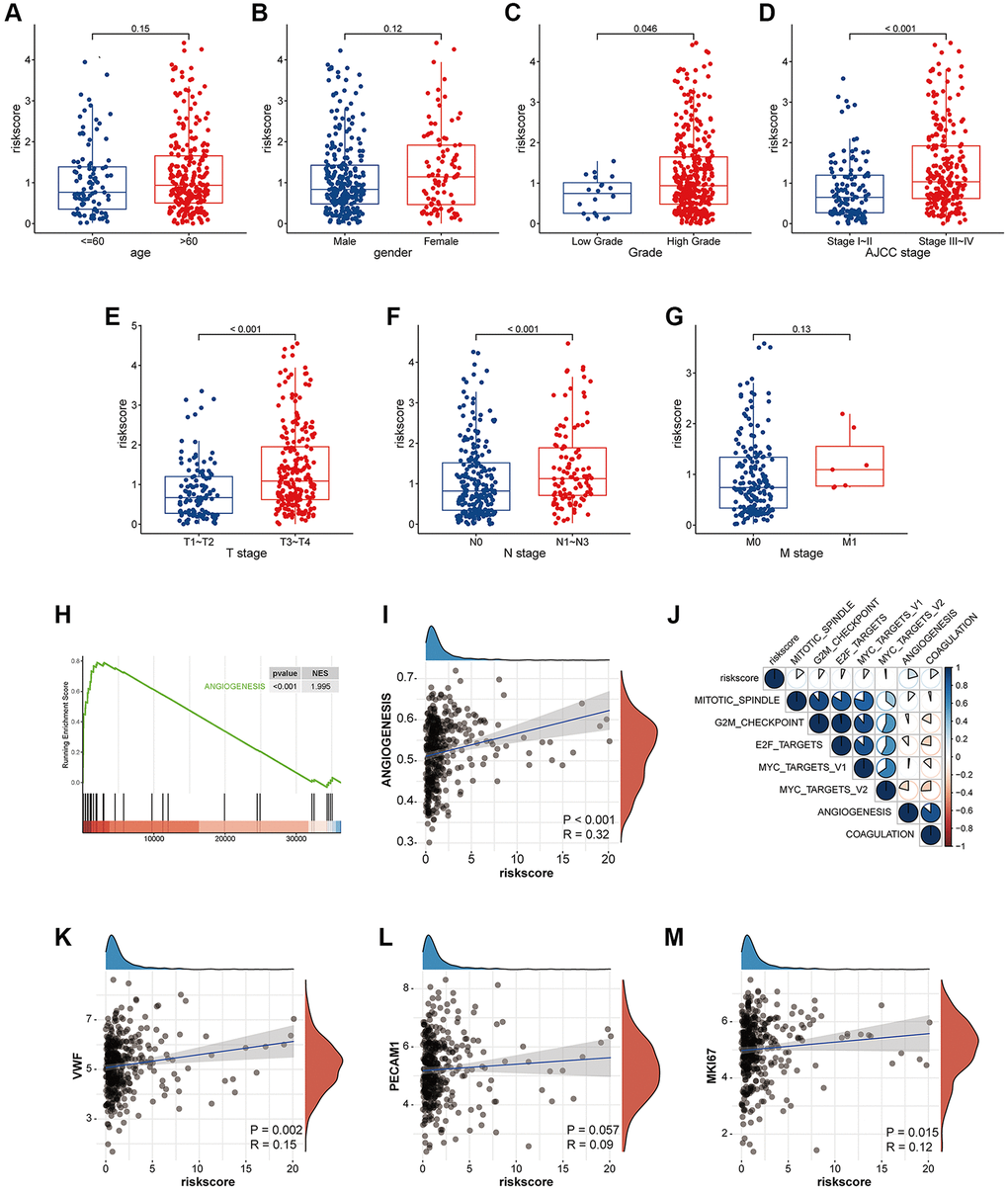
Correspondingly, the changes of coagulation state in the body can also affect the biological function of tumors. Coagulation factors are believed to promote cancer invasion and metastasis by promoting the interaction of tumor cell migration and angiogenesis [28]. Neutrophil extracellular traps (NETs) promote tumor cell migration, angiogenesis, and hypercoagulable state [29]. However, whether the abnormal expression of coagulation-related genes affects the biological behavior of tumors is not fully elucidated, and the research on coagulation-related genes in bladder tumors has not been reported yet. In this study, we confirmed the clinical association of coagulation-related genes within tumors with bladder cancer. In large cohort studies, we found that abnormal expression of tumor-associated genes is closely related to the prognosis, angiogenesis, and tumor proliferation of bladder cancer.
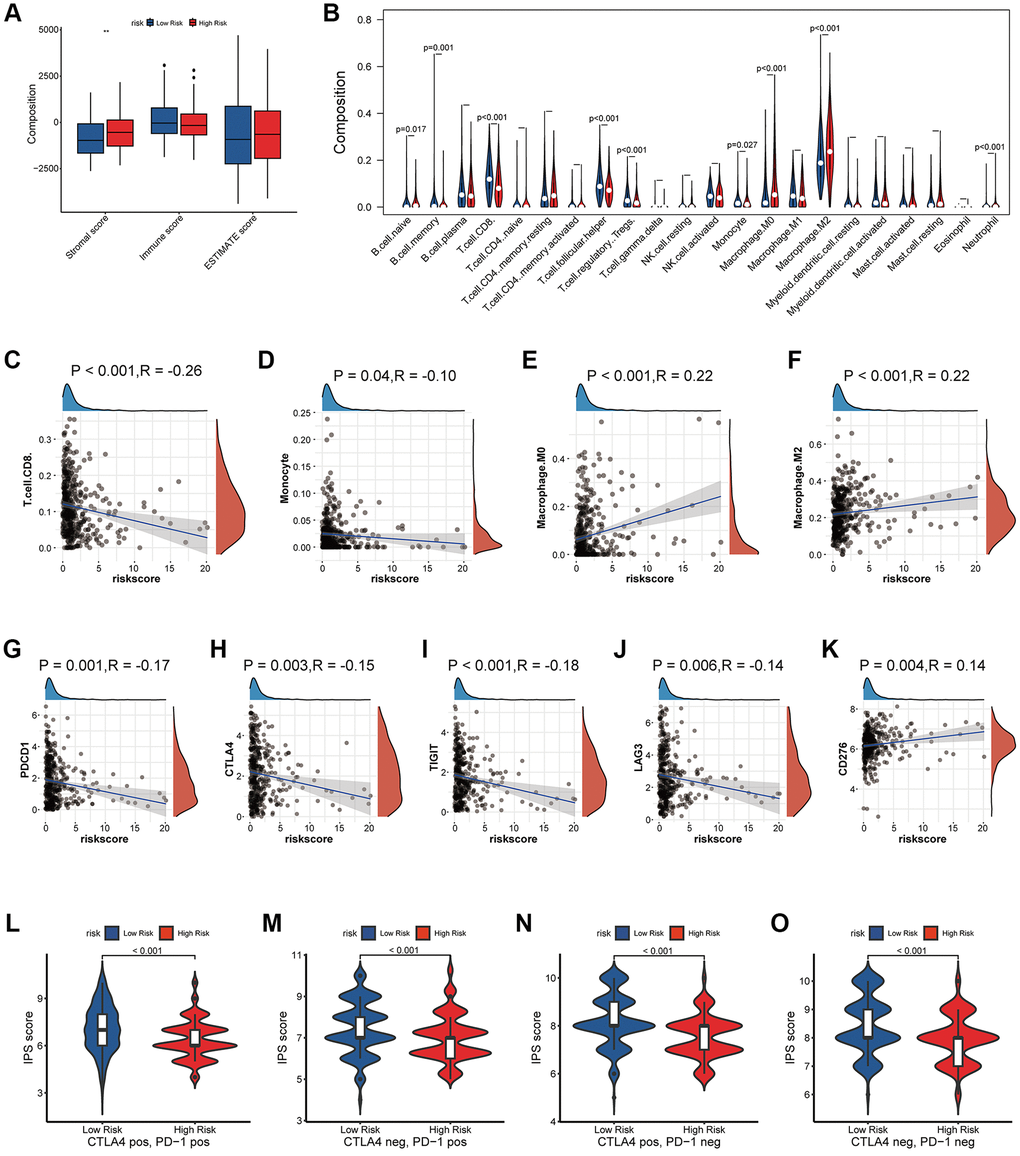
Pathway enrichment score indicated that coagulation genes not only activated procoagulant pathways, but also affected angiogenesis, tumor proliferation, and EMT pathways. Tumors need a large amount of energy in the process of growth and invasion, which can be accelerated by angiogenesis and remodeling. The need for angiogenesis in tumor growth and metastasis has been widely recognized [30]. As a symbolic behavior of malignant transformation of cells, EMT has also been widely studied [31]. In addition, our results also suggest that the abnormal expression of coagulation genes may be related to the inhibitory state of the immune microenvironment of bladder cancer. In our study, when the expression of coagulation-related genes was abnormal, immune-related pathways such as TGFβ-β, WNT-β-catenin, and IL6-STAT were all activated. The immune infiltration analysis indicated that when the coagulation related risk score was high, the infiltration abundance of B cells and T cells was low, while that of M2 macrophages was high. In our study, Regulatory T cells (Tregs) were more abundant in the low-risk group. Although considered to be an immunosuppressive cell population, the prognostic role of Tregs varies among tumor types [32]. In bladder cancer, there is a strong positive correlation between Tregs enrichment and T cell infiltration, higher infiltration abundance of Tregs indicates stronger immune cell enrichment and better prognosis [33, 34]. This is consistent with our research results. The state of the immune microenvironment is closely related to the therapeutic efficacy of PD1/PDL1. For example, when the WNT pathway is activated, even highly immunogenic tumor cells will still produce immune escape, thus affecting the therapeutic efficacy of PD1 inhibitors [35]. TGFβ can affect the sensitivity of tumor cells to cytotoxic T cells through a variety of pathways [36, 37]. In our present study, abnormal expression of coagulation genes affected the efficacy of PD1/PDL1 inhibitors for bladder cancer. A high risk associated with coagulation predicts a worse prognosis and a lower response rate.
As a retrospective study, we demonstrated the efficacy of the coagulation-related risk signature in different cohorts, but confounders that could influence clinical outcomes could not be excluded. Although our results confirm that the coagulation score is relevant to the prognosis of bladder cancer and the therapeutic efficacy of PD1/PDL1, further studies are needed to confirm the clinical utility of the coagulation score and its utility as a useful biomarker.
In conclusion, we provide a method for quantitative assessment of coagulation status within a tumor. At the same time, coagulation scores were strongly associated with angiogenesis, tumor proliferation, EMT, and poor prognosis in patients with bladder cancer as well as in those treated with PD1/PDL1.
Supplementary Materials
Author Contributions
ZL and YT designed the study; ZL, BL, YH, QL analyzed the data and wrote the manuscript; BL and QL contributed to draft editing; All authors have read and approved the final version of the manuscript.
Acknowledgments
We would like to express our appreciation to GEO, TCGA, UCSC Xena for providing the data used in this research study.
Conflicts of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The fund project name is Guangdong Medical Science and Technology Research Fund, and the project number is B2022027.
Editorial Note
This corresponding author has a verified history of publications using the personal email address for correspondence.
References
- 1. Kim AS, Khorana AA, McCrae KR. Mechanisms and biomarkers of cancer-associated thrombosis. Transl Res. 2020; 225:33–53. https://doi.org/10.1016/j.trsl.2020.06.012 [PubMed]
- 2. Labelle M, Begum S, Hynes RO. Direct signaling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell. 2011; 20:576–90. https://doi.org/10.1016/j.ccr.2011.09.009 [PubMed]
- 3. Parisi R, Panzera T, Russo L, Gamba S, De Curtis A, Di Castelnuovo A, Marchetti M, Cerletti C, Falanga A, de Gaetano G, Donati MB, Iacoviello L, Costanzo S, and Moli-sani Study Investigators. Fibrinogen levels in relation to colorectal cancer onset: A nested case-cohort study from the Moli-sani cohort. Front Cardiovasc Med. 2022; 9:1009926. https://doi.org/10.3389/fcvm.2022.1009926 [PubMed]
- 4. Henrikson KP, Salazar SL, Fenton JW
2nd , Pentecost BT. Role of thrombin receptor in breast cancer invasiveness. Br J Cancer. 1999; 79:401–6. https://doi.org/10.1038/sj.bjc.6690063 [PubMed] - 5. Diao X, Guo C, Li S. Identification of a novel anoikis-related gene signature to predict prognosis and tumor microenvironment in lung adenocarcinoma. Thorac Cancer. 2023; 14:320–30. https://doi.org/10.1111/1759-7714.14766 [PubMed]
- 6. Wang Y, Lin MG, Meng L, Chen ZM, Wei ZJ, Ying SC, Xu A. Identification of necroptosis-related genes for predicting prognosis and exploring immune infiltration landscape in colon adenocarcinoma. Front Oncol. 2022; 12:941156. https://doi.org/10.3389/fonc.2022.941156 [PubMed]
- 7. Xu R, Wu X, Du A, Zhao Q, Huang H. Identification of cuproptosis-related long non-coding ribonucleic acid signature as a novel prognosis model for colon cancer. Am J Cancer Res. 2022; 12:5241–54. [PubMed]
- 8. Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, Sabedot TS, Malta TM, Pagnotta SM, Castiglioni I, Ceccarelli M, Bontempi G, Noushmehr H. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016; 44:e71. https://doi.org/10.1093/nar/gkv1507 [PubMed]
- 9. Lee JS, Leem SH, Lee SY, Kim SC, Park ES, Kim SB, Kim SK, Kim YJ, Kim WJ, Chu IS. Expression signature of E2F1 and its associated genes predict superficial to invasive progression of bladder tumors. J Clin Oncol. 2010; 28:2660–7. https://doi.org/10.1200/JCO.2009.25.0977 [PubMed]
- 10. Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EE III, Koeppen H, Astarita JL, Cubas R, Jhunjhunwala S, Banchereau R, Yang Y, et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018; 554:544–8. https://doi.org/10.1038/nature25501 [PubMed]
- 11. Rose TL, Weir WH, Mayhew GM, Shibata Y, Eulitt P, Uronis JM, Zhou M, Nielsen M, Smith AB, Woods M, Hayward MC, Salazar AH, Milowsky MI, et al. Fibroblast growth factor receptor 3 alterations and response to immune checkpoint inhibition in metastatic urothelial cancer: a real world experience. Br J Cancer. 2021; 125:1251–60. https://doi.org/10.1038/s41416-021-01488-6 [PubMed]
- 12. Wagner GP, Kin K, Lynch VJ. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci. 2012; 131:281–5. https://doi.org/10.1007/s12064-012-0162-3 [PubMed]
- 13. Monti S, Tamayo P, Mesirov J, Golub T. Consensus Clustering: A Resampling-Based Method for Class Discovery and Visualization of Gene Expression Microarray Data. Mach Learn. 2003; 52:91–118. https://doi.org/10.1023/A:1023949509487
- 14. Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015; 43:e47. https://doi.org/10.1093/nar/gkv007 [PubMed]
- 15. Friedman J, Hastie T, Tibshirani R. Regularization Paths for Generalized Linear Models via Coordinate Descent. J Stat Softw. 2010; 33:1–22. [PubMed]
- 16. Barros AJ, Hirakata VN. Alternatives for logistic regression in cross-sectional studies: an empirical comparison of models that directly estimate the prevalence ratio. BMC Med Res Methodol. 2003; 3:21. https://doi.org/10.1186/1471-2288-3-21 [PubMed]
- 17. Huang ML, Hung YH, Lee WM, Li RK, Jiang BR. SVM-RFE based feature selection and Taguchi parameters optimization for multiclass SVM classifier. ScientificWorldJournal. 2014; 2014:795624. https://doi.org/10.1155/2014/795624 [PubMed]
- 18. Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, Müller M. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics. 2011; 12:77. https://doi.org/10.1186/1471-2105-12-77 [PubMed]
- 19. Zhou RS, Zhang EX, Sun QF, Ye ZJ, Liu JW, Zhou DH, Tang Y. Integrated analysis of lncRNA-miRNA-mRNA ceRNA network in squamous cell carcinoma of tongue. BMC Cancer. 2019; 19:779. https://doi.org/10.1186/s12885-019-5983-8 [PubMed]
- 20. Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015; 12:453–7. https://doi.org/10.1038/nmeth.3337 [PubMed]
- 21. Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012; 16:284–7. https://doi.org/10.1089/omi.2011.0118 [PubMed]
- 22. Wang D, Ye Q, Gu H, Chen Z. The role of lipid metabolism in tumor immune microenvironment and potential therapeutic strategies. Front Oncol. 2022; 12:984560. https://doi.org/10.3389/fonc.2022.984560 [PubMed]
- 23. Elyamany G, Alzahrani AM, Bukhary E. Cancer-associated thrombosis: an overview. Clin Med Insights Oncol. 2014; 8:129–37. https://doi.org/10.4137/CMO.S18991 [PubMed]
- 24. Donnellan E, Kevane B, Bird BR, Ainle FN. Cancer and venous thromboembolic disease: from molecular mechanisms to clinical management. Curr Oncol. 2014; 21:134–43. https://doi.org/10.3747/co.21.1864 [PubMed]
- 25. Lee AY. Management of thrombosis in cancer: primary prevention and secondary prophylaxis. Br J Haematol. 2005; 128:291–302. https://doi.org/10.1111/j.1365-2141.2004.05292.x [PubMed]
- 26. Falanga A, Gordon SG. Isolation and characterization of cancer procoagulant: a cysteine proteinase from malignant tissue. Biochemistry. 1985; 24:5558–67. https://doi.org/10.1021/bi00341a041 [PubMed]
- 27. Bick RL. Cancer-associated thrombosis. N Engl J Med. 2003; 349:109–11. https://doi.org/10.1056/NEJMp030086 [PubMed]
- 28. Wojtukiewicz MZ, Hempel D, Sierko E, Tucker SC, Honn KV. Protease-activated receptors (PARs)--biology and role in cancer invasion and metastasis. Cancer Metastasis Rev. 2015; 34:775–96. https://doi.org/10.1007/s10555-015-9599-4 [PubMed]
- 29. Efrimescu CI, Buggy PM, Buggy DJ. Neutrophil Extracellular Trapping Role in Cancer, Metastases, and Cancer-Related Thrombosis: a Narrative Review of the Current Evidence Base. Curr Oncol Rep. 2021; 23:118. https://doi.org/10.1007/s11912-021-01103-0 [PubMed]
- 30. Katsuta E, Rashid OM, Takabe K. Clinical relevance of tumor microenvironment: immune cells, vessels, and mouse models. Hum Cell. 2020; 33:930–7. https://doi.org/10.1007/s13577-020-00380-4 [PubMed]
- 31. Pastushenko I, Blanpain C. EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol. 2019; 29:212–26. https://doi.org/10.1016/j.tcb.2018.12.001 [PubMed]
- 32. Shang B, Liu Y, Jiang SJ, Liu Y. Prognostic value of tumor-infiltrating FoxP3+ regulatory T cells in cancers: a systematic review and meta-analysis. Sci Rep. 2015; 5:15179. https://doi.org/10.1038/srep15179 [PubMed]
- 33. Leblond MM, Zdimerova H, Desponds E, Verdeil G. Tumor-Associated Macrophages in Bladder Cancer: Biological Role, Impact on Therapeutic Response and Perspectives for Immunotherapy. Cancers (Basel). 2021; 13:4712. https://doi.org/10.3390/cancers13184712 [PubMed]
- 34. Koll FJ, Banek S, Kluth L, Köllermann J, Bankov K, Chun FK, Wild PJ, Weigert A, Reis H. Tumor-associated macrophages and Tregs influence and represent immune cell infiltration of muscle-invasive bladder cancer and predict prognosis. J Transl Med. 2023; 21:124. https://doi.org/10.1186/s12967-023-03949-3 [PubMed]
- 35. Takeuchi Y, Tanegashima T, Sato E, Irie T, Sai A, Itahashi K, Kumagai S, Tada Y, Togashi Y, Koyama S, Akbay EA, Karasaki T, Kataoka K, et al. Highly immunogenic cancer cells require activation of the WNT pathway for immunological escape. Sci Immunol. 2021; 6:eabc6424. https://doi.org/10.1126/sciimmunol.abc6424 [PubMed]
- 36. Bagati A, Kumar S, Jiang P, Pyrdol J, Zou AE, Godicelj A, Mathewson ND, Cartwright ANR, Cejas P, Brown M, Giobbie-Hurder A, Dillon D, Agudo J, et al. Integrin αvβ6-TGFβ-SOX4 Pathway Drives Immune Evasion in Triple-Negative Breast Cancer. Cancer Cell. 2021; 39:54–67.e9. https://doi.org/10.1016/j.ccell.2020.12.001 [PubMed]
- 37. Larson C, Oronsky B, Carter CA, Oronsky A, Knox SJ, Sher D, Reid TR. TGF-beta: a master immune regulator. Expert Opin Ther Targets. 2020; 24:427–38. https://doi.org/10.1080/14728222.2020.1744568 [PubMed]