Introduction
Renal cell carcinoma (RCC) is an intricate malignancy that originates from epithelial cells, with renal clear cell carcinoma (ccRCC) being the most common subtype [1]. The incidence and mortality of ccRCC have been steadily increasing, now accounting for approximately two to three percent of adult malignancies [2, 3]. Given its insensitivity to targeted and immunosuppressive agents, surgical intervention remains the primary and most effective treatment modality [4]. Despite significant advancements in early screening and diagnosis, approximately 30% of patients present with metastases at the time of diagnosis and about 25% develop metastases following surgical treatment [5, 6]. Consequently, the urgency lies in the quest for an effective prognostic signature and potential biomarkers to enhance the treatment of ccRCC patients.
RCC represents a prototypical immunogenic tumor that predominantly relies on inducing immunosuppressive cells, such as regulatory T cells, myeloid-derived suppressor cells, and macrophages, to create an immunosuppressive microenvironment [7]. The immune microenvironment has a dual role—it can inhibit tumor growth but also facilitate tumor progression by altering tumor immunogenicity or immunosuppression status [8]. Within the RCC tumor microenvironment, immunosuppressive cells may disrupt immune surveillance, ultimately leading to tumor immune evasion or escape [9]. RCC achieves this by upregulating the expression of immunosuppressive checkpoints, thereby inhibiting the activity of effector T cells and antigen-presenting cells, thus promoting tumor metastasis [10]. Consequently, it becomes imperative to explore the immune-related prognostic model for ccRCC.
Long noncoding RNAs (lncRNAs) are RNA transcripts exceeding 200 nucleotides in length that do not encode proteins; they are distributed widely in both the cytosol and nucleus [11, 12]. A growing body of evidence underscores the pivotal role of lncRNAs in regulating gene expression, translation and tumor progression [13, 14]. eRNAs are a class of RNA transcribed from enhancer regions on the genome, and they are found abundantly in most human cells and tissues [15, 16]. There is mounting evidence linking eRNA transcriptional levels to enhancer activity, implicating them in gene transcriptional regulation and their close association with tumor proliferation and metastasis [17, 18]. Moreover, eRNAs may contribute to tumor progression by regulating nuclear histone structure or interacting with transcriptional regulators [19, 20]. In human cells, eRNA participate in various signal transduction pathways and influence the construction of immune microenvironment by mediating the activation of target genes, thus underscoring the clinical significance of eRNA-targeted therapy [17]. Although immune-related eRNAs (IREs) play a substantial role in gene transcriptional control, their underlying mechanisms in ccRCC remain elusive.
In this study, we conducted a comprehensive evaluation of the prognostic characteristics of IREs in ccRCC. Notably, we established two distinct clusters of IREs, each with unique prognostic and immune characteristics. Additionally, we developed an IREs prognostic model that effectively predicts the survival rates of ccRCC patients. AC003092.1, a key player in the regulation of the tumor immune microenvironment, holds promise in guiding the development of immunotherapies for ccRCC.
Methods
Clinical data acquisition and extraction
Gene expression profiles and clinical data of ccRCC patients were sourced from The Cancer Genome Atlas (TCGA). The dataset comprised 538 cases of ccRCC tissues and 72 cases of normal tissues. For validation, gene expression data for ccRCC were obtained from ArrayExpress (https://www.ebi.ac.uk/arrayexpress) and the International Cancer Genome Consortium (ICGC) (https://icgc.org). The ArrayExpress dataset has the accession number E-MTAB-1980, including 106 cases with follow-up information. The ICGC dataset is labeled as RECA-EU and encompasses 91 cases with follow-up data.
Identification of the key eRNA in ccRCC
Based on the significant IREs expression profiles identified through univariate Cox analysis, patients were divided into high and low expression groups, and survival characteristics in ccRCC were analyzed further. Co-expression analysis was used to assess the correlation between IREs expression levels and their predicted target genes. IREs were included if they demonstrated a significant association with overall survival (OS) (KM log rank p < 0.05) and a significant association with predicted target genes (|r| >0.4 and p < 0.001). Key IREs, most relevant for survival according to log rank p-values, were selected for further analysis. The prognostic features of key IREs and their correlation with clinicopathological features were also investigated.
Cell culture and plasmid construction
We obtained two human ccRCC cell lines (Caki-1 and 769-P) and a human renal proximal tubular epithelial HK2 cell line (HRPTEpiC) that were purchased from the cell bank of the Chinese Academy of Sciences (Shanghai, China). All cells were cultured in RPMI 1640 medium (Thermo Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Thermo Fisher Scientific, Inc.) at a constant temperature of 37°C in a humidified atmosphere containing 5% CO2.
To silence AC003092.1, two siRNAs were transfected into Caki-1 and 769-P cells using Lipofectamine 3000 (Thermo Fisher Scientific, Inc.), following the manufacturer’s instructions. The sequences used for siRNA-1 were: Sense: GUAAUCCAGCGAAUCUGGA; Antisense: UCCAGAUUCGCUGGAUUAC; siRNA-2: Sense: CAGCAAUCAACAUAAUCAA; Antisense: UUGAUUAUGUUGAUUGCUG.
Cell counting kit-8 (CCK8) assay
Briefly, Caki-1 and 769-P cells, after various interventions, were incubated in 96-well plates (2 × 103) with 200 μL of culture medium at 37°C with 5% CO2. On days one, two, three, four and five, 20 μL CCK-8 solution was added into each well, and incubation was carried out for two hours. Absorbance was measured at an optical density of 450 nm using a Microplate reader (Bio-Rad Laboratories Inc., Hercules, CA, USA). Experiments were conducted in triplicate.
Cell growth assay and clonal formation assay
For the Cell Growth Assay, cell viability was assessed in accordance with the manufacturer’s instructions. Cells were initially seeded at a density of 2 × 103 cells per well in 96-well plates and evaluated at 0, 24, 48, 72, and 96 hours by the Cell Counting Kit-8 (Beyotime, Shanghai, China) and the Synergy H1 microplate reader (BioTek, Winooski, VT, USA) at 450 nm. In the case of the Clonal Formation Assay, after transfection and selection, 200 cells were distributed in 6-well plates in triplicate and incubated for 14 days. Subsequently, the cells were fixed with 10% ice-cold methanol and stained with 0.5% crystal violet solution. Colonies consisting more than 50 cells per colony were counted, and independent experiments were conducted in triplicate.
Data availability statement
All data used in this work can be acquired from (TCGA, (https://portal.gdc.cancer.gov/), GEO (https://www.ncbi.nlm.nih.gov/geo/).
Results
Identification of prognostic characteristics of the key eRNA---AC003092.1
As detailed in Supplementary Table 2, AC003092.1 emerged as the eRNA most strongly associated with survival, with a notably positive correlation to its predicted target, TFPI2. Subsequently, patients were evenly stratified into high and low AC003092.1 expression groups based on the median expression of AC003092.1 in ccRCC. Differences in survival outcomes, including OS, disease specific survival (DSS), and progress free interval (PFI) were assessed between these two groups. The analysis revealed that the high AC003092.1 group exhibited an adverse prognosis in OS, DSS, and PFI compared to the low AC003092.1 group, and these differences held statistical significance (Figure 5A–5C). Furthermore, when comparing AC003092.1 expression levels between ccRCC and adjacent tissues, AC003092.1 was found to be significantly upregulated in ccRCC (Figure 5D). The KM survival curve demonstrated that the high TFPI2 group was associated with a less favorable prognosis for ccRCC patients (Figure 5E). Additionally, we delved into the relationship between AC003092.1 expression and clinicopathological characteristics in ccRCC, revealing a significant positive correlation between the expression level of AC003092.1 and several clinicopathological features, including patient status, histological grade, pathological stage, and TNM stage (Figure 5F–5J). Furthermore, external validation sets, E-MTAB-1980 and ICGC, corroborated the association of AC003092.1 expression with a poor prognosis (Figure 5K, 5L).
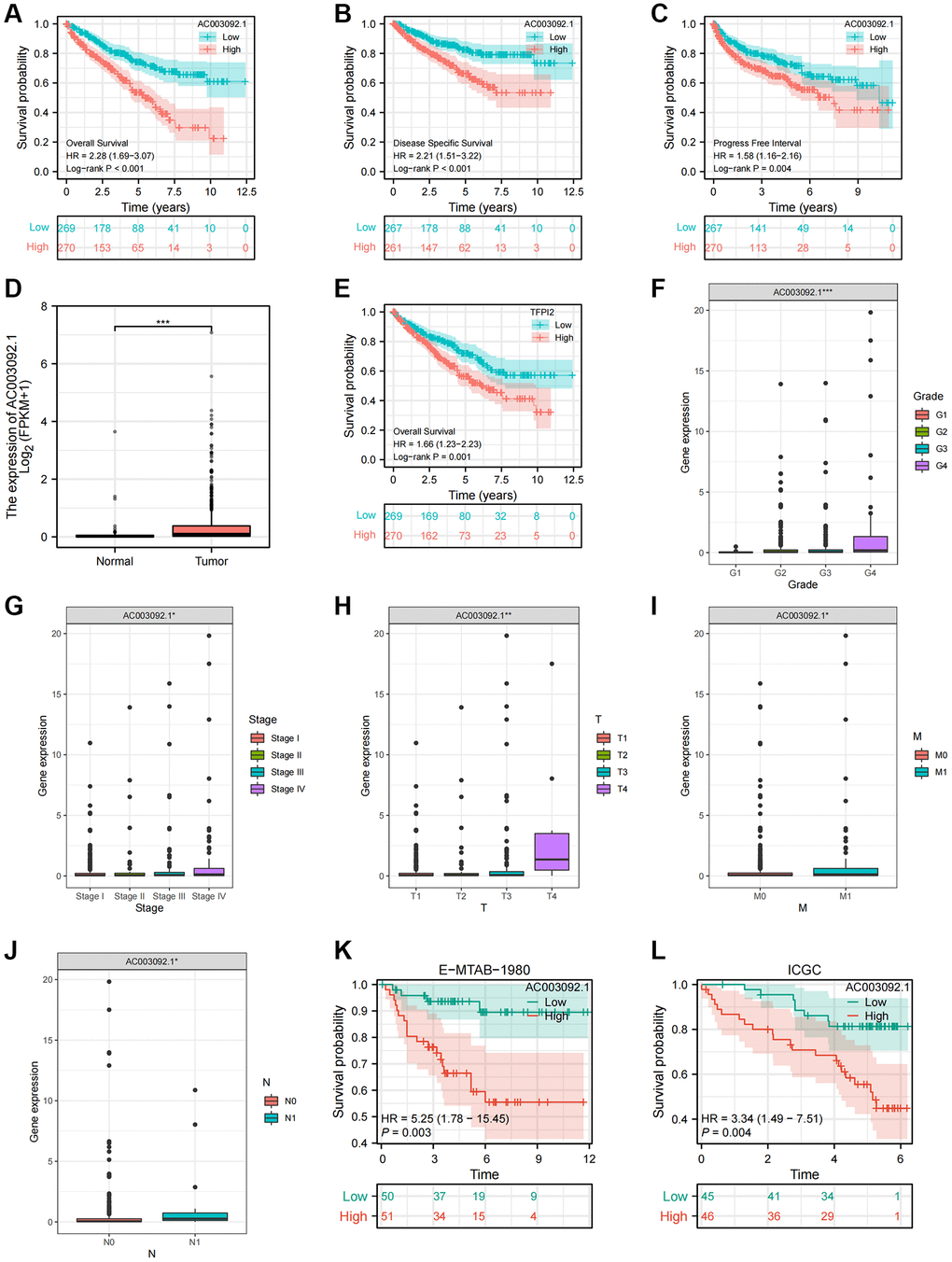
Figure 5. Clinical and prognostic characteristics of AC003092.1 in ccRCC. (A–C) KM survival curve showing the survival differences between high and low AC003092.1 groups (A: OS; B: DSS; C: PFI); (D) Difference of AC003092.1 mRNA expression between ccRCC and adjacent tissues; (E) KM survival curve showing the survival differences between high and low TFPI2 groups; (F–J) Differences of AC003092.1 expression in different clinicopathological variables (F: Grade; G: Stage; H: T stage; I: M stage; J: N stage); (K, L) KM survival curve showing the survival differences between high and low AC003092.1 groups in the E-MTAB-1980 and ICGC datasets.
Identification of immune characteristics of the key eRNA---AC003092.1
To gain deeper insights into the role of AC003092.1 in the tumor microenvironment, we explored the correlation between AC003092.1 and immune cells as well as tumor microenvironment scores. A heatmap highlighted that immunoinfiltrating cells and tumor microenvironment score were notably upregulated in the high AC003092.1 group (Figure 6A). Figure 6B presented the correlation between AC003092.1 and immune infiltrating cells, revealing a significantly positively correlation with immunosuppressive cells (MDSC, Macrophage and Regulatory.T.cell) and a negative correlation with NK cells and neutrophils. This analysis was complemented by a noticeable increase in the expression of immunosuppressive cells in the high AC003092.1 group when compared to the low AC003092.1 group (Figure 6C–6E). Additionally, the tumor microenvironment score, encompassing estimated score, immune score, and stromal score, was significantly elevated in the high AC003092.1 group (Figure 6F–6H). Further exploration of the correlation between AC003092.1 and immunosuppressive checkpoints revealed that immunosuppressive checkpoints (TGFBR1, CTLA4, and CD96) were also significantly upregulated in the high AC003092.1 group (Figure 6I–6K).
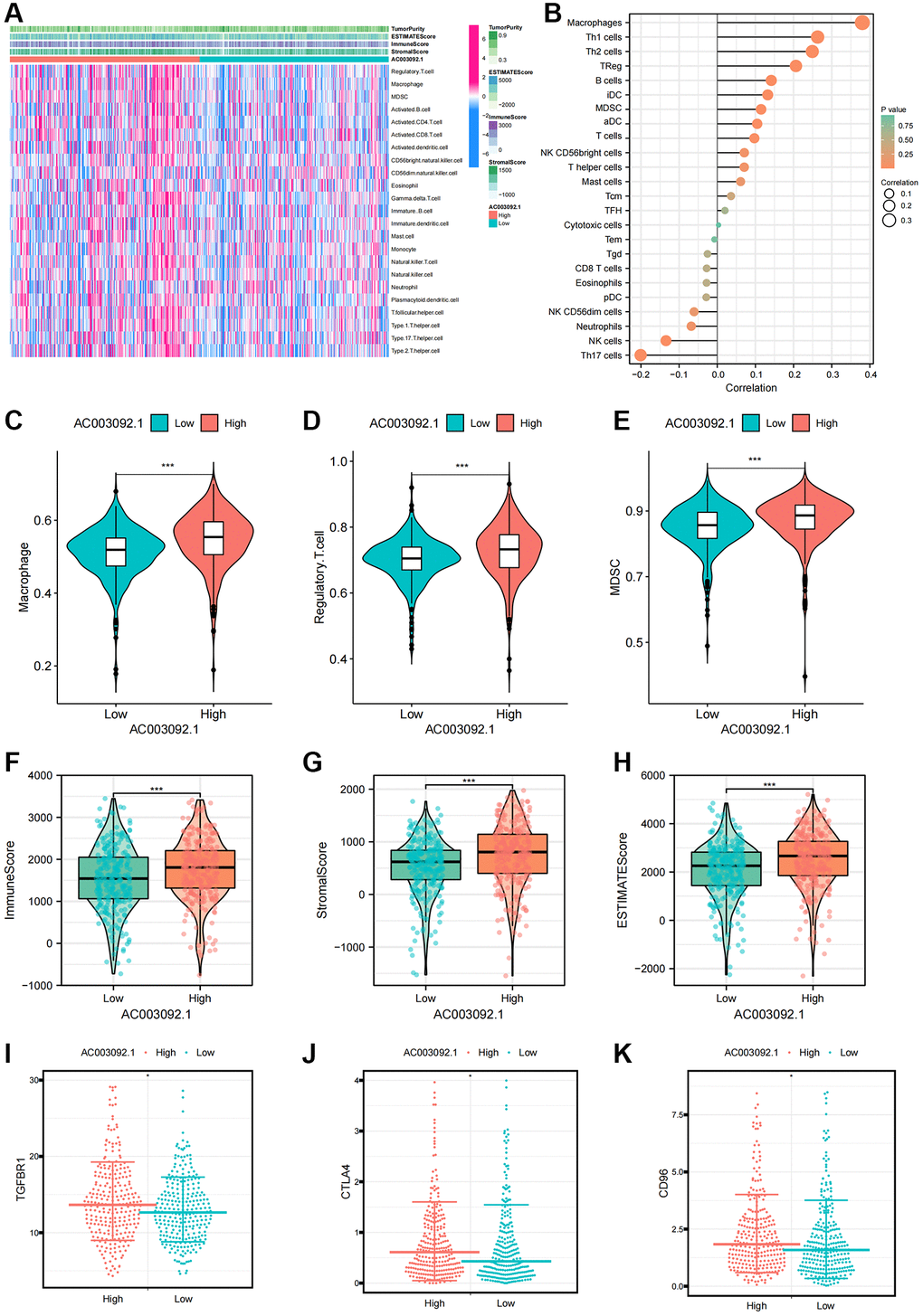
Figure 6. Immune characteristics of AC003092.1 in ccRCC. (A) Distribution of immune infiltrating cells and tumor microenvironment scores in high and low AC003092.1 expression groups; (B) Correlation between immunoinfiltrating cells and AC003092.1 expression profiles. (C–E) Differences in immunosuppressive cell expression between high and low AC003092.1 groups (C: Macrophage; D: Regulatory.T.cell; E: MDSC); (F–H) Differences in tumor microenvironment scores between high and low AC003092.1 groups (F: ImmuneScore; G: StromalScore; H: ESTIMATEScore); (I–K) Differences in immune suppression checkpoints between high and low AC003092.1 groups (I: TGFBR1; J: CTLA4; K: CD96).
AC003092-knockdown suppressed proliferation in Caki-1 and 769-P cells
RT-qPCR results indicated that AC003092.1 was substantially upregulated in ccRCC tissues (Figure 7A). Comparative analysis with HK2 cell lines showed a significant increase in AC003092.1 expression in renal cell lines, particularly in 769-P and Caki-1, with the highest expression observed in Caki-1 (Figure 7B). Subsequently, AC003092.1-shRNA was transfected into 769-P and Caki-1 cells to effectively knock down AC003092.1, confirmed by RT-qPCR (Figure 7C). Next, CCK8 assay revealed that AC003092.1 knockdown led to a reduction in proliferation in both 769-P and Caki-1 cells (Figure 7D). In the clonogenic assay, AC003092.1 knockdown resulted in a significantly lower number of colonies formed in both 769-P and Caki-1 cells compared to the empty vector controls (Figure 7E).
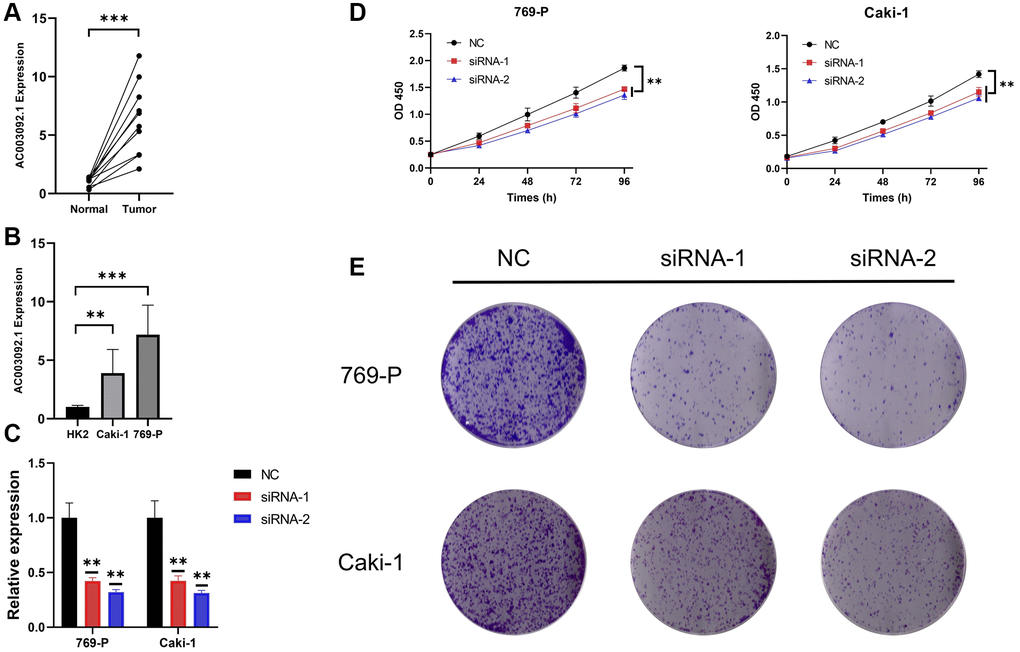
Figure 7. AC003092-knockdown suppressed proliferation in caki-1 and 769-P cells. (A) Differences in AC003092.1 expression between ccRCC tissues and adjacent tissues; (B) Differences in AC003092.1 expression between HK2 and renal cell lines 769-P and Caki-1; (C) The expression of AC003092 was downregulated in 769-P and Caki-1 cells, respectively, as determined by RT-qPCR; (D) AC003092-knockdown suppressed ccRCC cell proliferation in 769-P and Caki-1 cells; (E) Clonogenic formation in AC003092-knockdown 769-P and Caki-1 cells, along with images of formed clones following control cells for two weeks.
Discussion
Epidemiological studies have revealed a rising incidence and mortality rate of RCC. RCC is characterized as an immunogenic tumor with infiltrating myeloid cell, including macrophages and neutrophils as well as CD8+ T cells and natural killer (NK) cells [23]. Despite the identification of immune cells and immune checkpoints as novel prognostic biomarkers and therapeutic targets for ccRCC, only a fraction of patients with ccRCC benefit from such approaches [9, 24]. The dysregulation of eRNAs is closely linked to various human diseases and immune microenvironment, making eRNAs a promising target for effective therapeutic interventions [25, 26]. Therefore, our objective was to construct a novel IREs prognostic model for ccRCC, aiming to identify reliable predictive and prognostic biomarkers while exploring new immunotherapeutic targets.
eRNA represents a noncoding RNA transcribed by enhancers, facilitating the activation of target genes [27, 28]. These eRNAs not only play a role in regulating the immune response but are also involved in numerous tumorigenic signaling pathways, such as p53 and immune checkpoints, which hold pivotal roles in tumor progression and metastasis [29]. For instance, KLK3 eRNA (KLK3e) selectively enhances the expression of androgen receptor-regulated genes, thereby promoting the proliferation and metastasis of prostate cancer [30]. NET1e is significantly overexpressed in breast cancer and is associated with poor prognosis [17]. As our understanding of eRNA mechanisms deepens, studies have revealed that eRNAs fulfill diverse biological roles in the metastasis and progression of various tumors [31]. Fan et al. constructed an eRNA-related prognostic model in prostate cancer that effectively predicted patient outcomes and explored the immune characteristics of this model [32]. The similarity between our study and other model-based research lies in the fact that both involve clustering key genes, building models, and subsequently analyzing the model’s immune, mutation, and prognostic features [33–38]. Notably, our study extended beyond the prognosis and immune characteristics of the IREs signature; we also identified the most relevant eRNA for survival and conducted in-depth analyses of its immune and clinicopathological characteristics.
In our investigation, AC003092.1 emerged as the IRE most strongly associated with survival in ccRCC. AC003092.1 was significantly upregulated in ccRCC and exhibited a close association with poor prognosis and clinicopathological staging. Furthermore, AC003092.1 displayed a significant positive correlation with immunosuppressive cells and immunosuppressive checkpoints, suggesting its potential involvement in shaping an immunosuppressive microenvironment. Notably, Guo et al. demonstrated that AC003092.1 was an IRE, linked to immune cell composition, function, and pathways, possibly contributing to the formation of glioblastoma multiforme (GBM). In GBM patients, AC003092.1 was significantly correlated with poor prognosis and the upregulated expression of its target gene, TFPI2 [39]. AC003092.1’s role in GBM involves enhancing the sensitivity of GBM to temozolomide through the mediation of the miR-195/TFPI-2 signaling pathway, impacting patient prognosis. Furthermore, AC003092.1 can counteract the upregulation of TFPI2 expression by miR-195, thus promoting temozolomide-induced apoptosis [40]. These findings highlighted AC003092.1 as a promising molecular target for preventive and therapeutic strategies of ccRCC. However, a comprehensive understanding of the molecular mechanisms governing AC003092.1’s aberrant regulation and its role in ccRCC progression necessitates further investigation, ideally through in vivo and in vitro experimental analysis.
Nonetheless, it is important to acknowledge certain limitations in our study. Firstly, our data analysis relied on publicly available datasets, and additional datasets should be employed for further validation of our results. Additionally, the biological mechanism underlying AC003092.1’s influence on reshaping the immunosuppressive microenvironment warrant exploration through in-depth in vivo and in vitro experiments.
Conclusion
IREs played a pivotal role in shaping the immunosuppressive TME in ccRCC. The IREs signature demonstrated remarkable accuracy in distinguishing the immune characteristics and predicting the prognosis of ccRCC patients. AC003092.1, specifically, exhibited an immunosuppressive effect within the TME and hold promise as a potential therapeutic target for ccRCC treatment.
Author Contributions
Yang Li and Hongqi Chen designed this work. Lili Niu and Yulin Yue were responsible for the revised and funding. Qiang Li wrote the manuscript. Yang Lv performed the bioinformatics analysis and performed experiments. Wenchuan Shao and Xinghan Yan performed the data review. All authors have read and approved the manuscript.
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
Ethical Statement
The protocols for cell experiments adhered to established ethical guidelines and standards for cellular research. We understand the importance of ethical considerations in scientific research and confirm that our study was conducted in accordance with the relevant ethical guidelines.
Funding
This research was supported by the Suzhou Science and Technology Plan Project (No. SKY2022032, No. SKYD2023023 and SKJYD2021030), “Science, Education, and Health Promotion” Project in Wujiang District, Suzhou (No. wwk201717 and wwk202104).
References
- 1. Shuch B, Amin A, Armstrong AJ, Eble JN, Ficarra V, Lopez-Beltran A, Martignoni G, Rini BI, Kutikov A. Understanding pathologic variants of renal cell carcinoma: distilling therapeutic opportunities from biologic complexity. Eur Urol. 2015; 67:85–97. https://doi.org/10.1016/j.eururo.2014.04.029 [PubMed]
- 2. Barata PC, Rini BI. Treatment of renal cell carcinoma: Current status and future directions. CA Cancer J Clin. 2017; 67:507–24. https://doi.org/10.3322/caac.21411 [PubMed]
- 3. Ljungberg B, Bensalah K, Canfield S, Dabestani S, Hofmann F, Hora M, Kuczyk MA, Lam T, Marconi L, Merseburger AS, Mulders P, Powles T, Staehler M, et al. EAU guidelines on renal cell carcinoma: 2014 update. Eur Urol. 2015; 67:913–24. https://doi.org/10.1016/j.eururo.2015.01.005 [PubMed]
- 4. Escudier B, Porta C, Schmidinger M, Rioux-Leclercq N, Bex A, Khoo V, Grünwald V, Gillessen S, Horwich A, and ESMO Guidelines Committee. Renal cell carcinoma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up†. Ann Oncol. 2019; 30:706–20. https://doi.org/10.1093/annonc/mdz056 [PubMed]
- 5. Hsieh JJ, Purdue MP, Signoretti S, Swanton C, Albiges L, Schmidinger M, Heng DY, Larkin J, Ficarra V. Renal cell carcinoma. Nat Rev Dis Primers. 2017; 3:17009. https://doi.org/10.1038/nrdp.2017.9 [PubMed]
- 6. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492 [PubMed]
- 7. Díaz-Montero CM, Rini BI, Finke JH. The immunology of renal cell carcinoma. Nat Rev Nephrol. 2020; 16:721–35. https://doi.org/10.1038/s41581-020-0316-3 [PubMed]
- 8. Giraldo NA, Becht E, Vano Y, Petitprez F, Lacroix L, Validire P, Sanchez-Salas R, Ingels A, Oudard S, Moatti A, Buttard B, Bourass S, Germain C, et al. Tumor-Infiltrating and Peripheral Blood T-cell Immunophenotypes Predict Early Relapse in Localized Clear Cell Renal Cell Carcinoma. Clin Cancer Res. 2017; 23:4416–28. https://doi.org/10.1158/1078-0432.CCR-16-2848 [PubMed]
- 9. Blankenstein T, Coulie PG, Gilboa E, Jaffee EM. The determinants of tumour immunogenicity. Nat Rev Cancer. 2012; 12:307–13. https://doi.org/10.1038/nrc3246 [PubMed]
- 10. Cella D, Grünwald V, Escudier B, Hammers HJ, George S, Nathan P, Grimm MO, Rini BI, Doan J, Ivanescu C, Paty J, Mekan S, Motzer RJ. Patient-reported outcomes of patients with advanced renal cell carcinoma treated with nivolumab plus ipilimumab versus sunitinib (CheckMate 214): a randomised, phase 3 trial. Lancet Oncol. 2019; 20:297–310. https://doi.org/10.1016/S1470-2045(18)30778-2 [PubMed]
- 11. Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012; 482:339–46. https://doi.org/10.1038/nature10887 [PubMed]
- 12. Lin W, Zhou Q, Wang CQ, Zhu L, Bi C, Zhang S, Wang X, Jin H. LncRNAs regulate metabolism in cancer. Int J Biol Sci. 2020; 16:1194–206. https://doi.org/10.7150/ijbs.40769 [PubMed]
- 13. Liang Y, Chen X, Wu Y, Li J, Zhang S, Wang K, Guan X, Yang K, Bai Y. LncRNA CASC9 promotes esophageal squamous cell carcinoma metastasis through upregulating LAMC2 expression by interacting with the CREB-binding protein. Cell Death Differ. 2018; 25:1980–95. https://doi.org/10.1038/s41418-018-0084-9 [PubMed]
- 14. Bhan A, Soleimani M, Mandal SS. Long Noncoding RNA and Cancer: A New Paradigm. Cancer Res. 2017; 77:3965–81. https://doi.org/10.1158/0008-5472.CAN-16-2634 [PubMed]
- 15. Li W, Notani D, Rosenfeld MG. Enhancers as non-coding RNA transcription units: recent insights and future perspectives. Nat Rev Genet. 2016; 17:207–23. https://doi.org/10.1038/nrg.2016.4 [PubMed]
- 16. Huang J, Wang J, He H, Huang Z, Wu S, Chen C, Liu W, Xie L, Tao Y, Cong L, Jiang Y. Close interactions between lncRNAs, lipid metabolism and ferroptosis in cancer. Int J Biol Sci. 2021; 17:4493–513. https://doi.org/10.7150/ijbs.66181 [PubMed]
- 17. Zhang Z, Lee JH, Ruan H, Ye Y, Krakowiak J, Hu Q, Xiang Y, Gong J, Zhou B, Wang L, Lin C, Diao L, Mills GB, et al. Transcriptional landscape and clinical utility of enhancer RNAs for eRNA-targeted therapy in cancer. Nat Commun. 2019; 10:4562. https://doi.org/10.1038/s41467-019-12543-5 [PubMed]
- 18. Peng WX, Koirala P, Mo YY. LncRNA-mediated regulation of cell signaling in cancer. Oncogene. 2017; 36:5661–7. https://doi.org/10.1038/onc.2017.184 [PubMed]
- 19. Tan SH, Leong WZ, Ngoc PCT, Tan TK, Bertulfo FC, Lim MC, An O, Li Z, Yeoh AEJ, Fullwood MJ, Tenen DG, Sanda T. The enhancer RNA ARIEL activates the oncogenic transcriptional program in T-cell acute lymphoblastic leukemia. Blood. 2019; 134:239–51. https://doi.org/10.1182/blood.2018874503 [PubMed]
- 20. Napoli S, Munz N, Guidetti F, Bertoni F. Enhancer RNAs (eRNAs) in Cancer: The Jacks of All Trades. Cancers (Basel). 2022; 14:1978. https://doi.org/10.3390/cancers14081978 [PubMed]
- 21. Murakami S, Gadad SS, Kraus WL. A PreSTIGEous use of LncRNAs to predict enhancers. Cell Cycle. 2015; 14:1619–20. https://doi.org/10.1080/15384101.2015.1032650 [PubMed]
- 22. Gu X, Wang L, Boldrup L, Coates PJ, Fahraeus R, Sgaramella N, Wilms T, Nylander K. AP001056.1, A Prognosis-Related Enhancer RNA in Squamous Cell Carcinoma of the Head and Neck. Cancers (Basel). 2019; 11:347. https://doi.org/10.3390/cancers11030347 [PubMed]
- 23. Komohara Y, Hasita H, Ohnishi K, Fujiwara Y, Suzu S, Eto M, Takeya M. Macrophage infiltration and its prognostic relevance in clear cell renal cell carcinoma. Cancer Sci. 2011; 102:1424–31. https://doi.org/10.1111/j.1349-7006.2011.01945.x [PubMed]
- 24. Nakano O, Sato M, Naito Y, Suzuki K, Orikasa S, Aizawa M, Suzuki Y, Shintaku I, Nagura H, Ohtani H. Proliferative activity of intratumoral CD8(+) T-lymphocytes as a prognostic factor in human renal cell carcinoma: clinicopathologic demonstration of antitumor immunity. Cancer Res. 2001; 61:5132–6. [PubMed]
- 25. Arnold PR, Wells AD, Li XC. Diversity and Emerging Roles of Enhancer RNA in Regulation of Gene Expression and Cell Fate. Front Cell Dev Biol. 2020; 7:377. https://doi.org/10.3389/fcell.2019.00377 [PubMed]
- 26. Léveillé N, Melo CA, Agami R. Enhancer-associated RNAs as therapeutic targets. Expert Opin Biol Ther. 2015; 15:723–34. https://doi.org/10.1517/14712598.2015.1029452 [PubMed]
- 27. Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S, Markenscoff-Papadimitriou E, Kuhl D, Bito H, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature. 2010; 465:182–7. https://doi.org/10.1038/nature09033 [PubMed]
- 28. Sartorelli V, Lauberth SM. Enhancer RNAs are an important regulatory layer of the epigenome. Nat Struct Mol Biol. 2020; 27:521–8. https://doi.org/10.1038/s41594-020-0446-0 [PubMed]
- 29. Melo CA, Drost J, Wijchers PJ, van de Werken H, de Wit E, Oude Vrielink JA, Elkon R, Melo SA, Léveillé N, Kalluri R, de Laat W, Agami R. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol Cell. 2013; 49:524–35. https://doi.org/10.1016/j.molcel.2012.11.021 [PubMed]
- 30. Hsieh CL, Fei T, Chen Y, Li T, Gao Y, Wang X, Sun T, Sweeney CJ, Lee GS, Chen S, Balk SP, Liu XS, Brown M, Kantoff PW. Enhancer RNAs participate in androgen receptor-driven looping that selectively enhances gene activation. Proc Natl Acad Sci U S A. 2014; 111:7319–24. https://doi.org/10.1073/pnas.1324151111 [PubMed]
- 31. Cai S, Hu X, Chen R, Zhang Y. Identification and Validation of an Immune-Related eRNA Prognostic Signature for Hepatocellular Carcinoma. Front Genet. 2021; 12:657051. https://doi.org/10.3389/fgene.2021.657051 [PubMed]
- 32. Fan S, Wang Z, Zhao L, Zhao C, Yuan D, Wang J. A Robust Prognostic Gene Signature Based on eRNAs-Driven Genes in Prostate Cancer. Front Genet. 2021; 12:676845. https://doi.org/10.3389/fgene.2021.676845 [PubMed]
- 33. Gao C, Zhang R, Chen X, Yao T, Song Q, Ye W, Li P, Wang Z, Yi D, Wu Y. Integrating Internet multisource big data to predict the occurrence and development of COVID-19 cryptic transmission. NPJ Digit Med. 2022; 5:161. https://doi.org/10.1038/s41746-022-00704-8 [PubMed]
- 34. Lai G, Zhong X, Liu H, Deng J, Li K, Xie B. Development of a Hallmark Pathway-Related Gene Signature Associated with Immune Response for Lower Grade Gliomas. Int J Mol Sci. 2022; 23:11971. https://doi.org/10.3390/ijms231911971 [PubMed]
- 35. Lai G, Zhong X, Liu H, Deng J, Li K, Xie B. A Novel m7G-Related Genes-Based Signature with Prognostic Value and Predictive Ability to Select Patients Responsive to Personalized Treatment Strategies in Bladder Cancer. Cancers (Basel). 2022; 14:5346. https://doi.org/10.3390/cancers14215346 [PubMed]
- 36. Lai G, Liu H, Deng J, Li K, Zhang C, Zhong X, Xie B. The Characteristics of Tumor Microenvironment Predict Survival and Response to Immunotherapy in Adrenocortical Carcinomas. Cells. 2023; 12:755. https://doi.org/10.3390/cells12050755 [PubMed]
- 37. Lai G, Liu H, Deng J, Li K, Xie B. A Novel 3-Gene Signature for Identifying COVID-19 Patients Based on Bioinformatics and Machine Learning. Genes (Basel). 2022; 13:1602. https://doi.org/10.3390/genes13091602 [PubMed]
- 38. Ma Q, Tao H, Li Q, Zhai Z, Zhang X, Lin Z, Kuang N, Pan J. OrganoidDB: a comprehensive organoid database for the multi-perspective exploration of bulk and single-cell transcriptomic profiles of organoids. Nucleic Acids Res. 2023; 51:D1086–93. https://doi.org/10.1093/nar/gkac942 [PubMed]
- 39. Guo XY, Zhong S, Wang ZN, Xie T, Duan H, Zhang JY, Zhang GH, Liang L, Cui R, Hu HR, Lu J, Wu Y, Dong JJ, et al. Immunogenomic Profiling Demonstrate AC003092.1 as an Immune-Related eRNA in Glioblastoma Multiforme. Front Genet. 2021; 12:633812. https://doi.org/10.3389/fgene.2021.633812 [PubMed]
- 40. Xu N, Liu B, Lian C, Doycheva DM, Fu Z, Liu Y, Zhou J, He Z, Yang Z, Huang Q, Zeng H, Guo H. Long noncoding RNA AC003092.1 promotes temozolomide chemosensitivity through miR-195/TFPI-2 signaling modulation in glioblastoma. Cell Death Dis. 2018; 9:1139. https://doi.org/10.1038/s41419-018-1183-8 [PubMed]