Introduction
Lung cancer is a highly malignant tumor that often leads to extremely high mortality. [1, 2]. Cancer stem cells (CSCs) have been extensively studied and proved to account for multidrug resistance and metastasis in various cancer types [3]. Therefore, mechanistic studies aiming to illustrate the molecular machinery required to initiate and maintain the malignant behavior of CSCs may pave the way for the final eradication of CSCs and improve the therapeutic efficacy of lung cancer.
Long noncoding RNAs play an indispensable function in the regulate the expression of oncogenes or tumor suppressor genes [4–7]. High-throughput next-generation sequencing has revealed that N6-methyladenine is the most abundant RNA modification in diverse cells and plays a pivotal function in the gene expression [8, 9]. m6A modification exerts its function via readers (IGF2BPs, METTL3/14) and erasers (FTO, ALKBH5) [10, 11]. However, information available on the expression, regulation, and clinical significance of m6A-related AC026356.1 in LUAD has remained lacking.
In this study, we investigated how IGF2BP2 forms a positive loop with the AC026356.1 signaling cascade through modulation at transcriptional and post-transcriptional levels. On one hand, IGF2BP2 maintains AC026356.1 transcription, acting as a positive transcription factor. On the other hand, AC026356.1 promoted the transcription of METTL14/IGF2BP2 and guided m6A modification. The m6A-modified AC026356.1 loop may serve as a potential prognostic predictor for lung cancer patients.
Materials and Methods
Data download
We obtained the RNA expression and clinical information of lung cancer from the TCGA-LUAD database (https://cancergenome.nih.gov/). We analysis of the expression, clinical information, and prognosis of AC026356.1 in lung cancer by using the above TCGA dataset.
LncACTdb database
LncACTdb is an updated database of experimentally supported ceRNA interactions. In this study, we use LncACTdb to analyze the potential AC026356.1-related ceRNA network [12].
Cell lines, cell culture, and qPCR assay
The lung cancer cell line was purchased from the cell bank of the ATCC and cultured in BEGM media (Lonza, CC-3170). The Qpcr primers as follows: AC026356.1-F: TAGAGAAAATTATTCTT; AC026356.1-R: GCTTGCCCTAACAGCAGC; AC026356.1-ASO: ATGTGAGTATAGGGCAGAT.
Cell proliferation assay
The cholecystokinin octapeptide (CCK-8) assay was performed 24, 48, 72, and 96 h after propagation. Cells were seeded and assessed in triplicates.
Tumor sphere formation assay
Cells (2 × 104) were seeded onto 12-well plates and cultured in serum-free 1640 medium. Cell spheroids were documented and quantified using an inverted microscope (Olympus, Japan) after two weeks.
Statistical analysis
Comparisons between groups were calculated using the Wilcoxon rank-sum or Kruskal-Wallis test. P < 0.05 was considered significant and marked with asterisks as indicated.
Data availability
All data used in the study were from The Cancer Genome Atlas (TCGA-LUAD) (https://portal.gdc.cancer.gov/).
Results
AC026356.1 is highly expressed in LUAD
We performed data mining using the TCGA dataset and confirmed the overexpression of AC026356.1 in lung cancer samples (Figure 1A, 1B). Lung cancer transcriptome sequencing data from Gene Expression Omnibus (GEO) also showed that AC026356.1 is highly expressed in lung cancer (Figure 1C). Elevation of AC026356.1 was positively correlated with the pathological stages (Figure 1D). We next determine the relationships between AC026356.1 expression and clinical feature. Results showed that forced expression of AC026356.1 was correlated with diverse survival events (Figure 1E, 1F). These results also validate by the GEO dataset (Figure 1G). While, AC026356.1 not affect the prognosis of lung squamous cell carcinoma (LUSC) patients (Figure 1H). ROC curve analysis results uncover that AC026356.1 may be a potential diagnostic marker in lung cancer (Figure 1I, 1J).
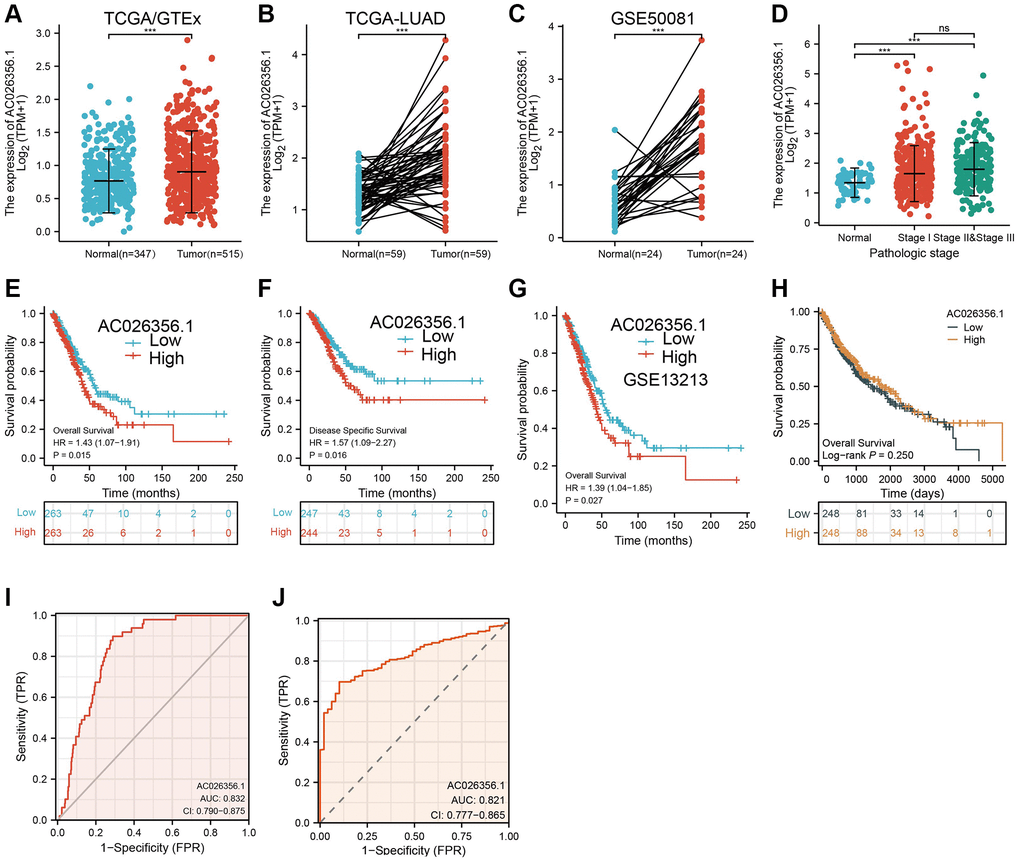
Figure 1. AC026356.1 was upregulated in LUAD. (A–C) AC026356.1 level in lung cancer by TCGA and GEO database. (D) Correlation between AC026356.1 expression and pathological stage in LUAD by TCGA database. (E–G) Prognosis of AC026356.1 in lung cancer examined by TCGA and GEO dataset. (H) Prognosis of AC026356.1 in LUSC examined by TCGA dataset. (I, J) Diagnostic value of AC026356.1 in NSCLC. NS > 0.05 and ***p < 0.001.
Analysis of subcellular localization and coding ability of AC026356.1
We further analysis of the genetic characteristics and secondary structure of AC026356.1 by using the UCSC database (http://genome.ucsc.edu/). As is shown in Figure 2A, 2B, AC026356.1 is located in chromosome 12 and has a complex spatial structure. Subcellular localization and coding ability analysis confirmed that AC026356.1 was predominantly resided in the cytoplasm of lung cancer cells and does not have protein-coding potential (Figure 2C–2F).
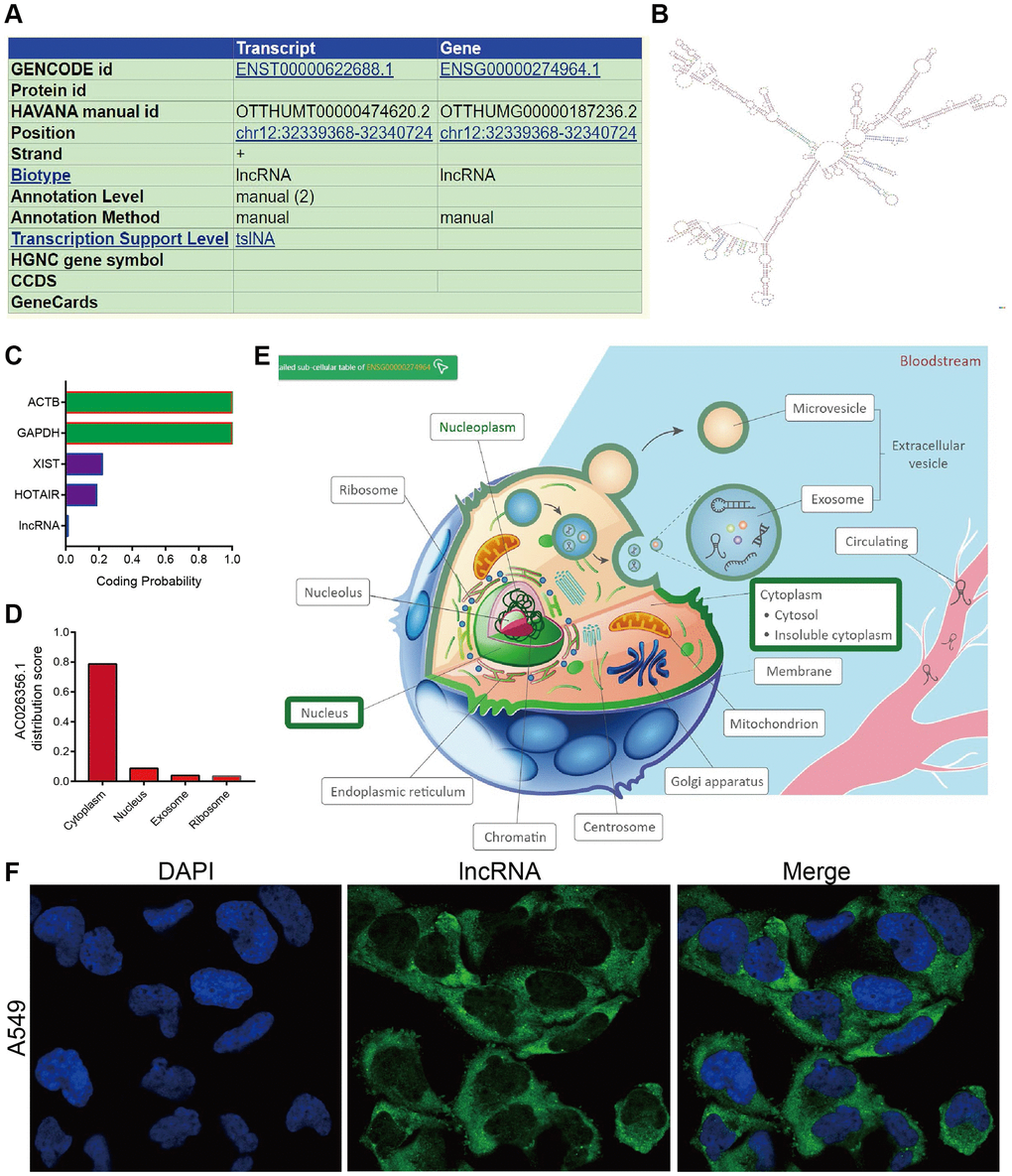
Figure 2. Gene characteristics analysis of AC026356.1. (A, B) The genetic characteristics and secondary structure of AC026356.1 by using the UCSC database (http://genome.ucsc.edu/). (C–E) Subcellular localization and coding ability analysis of AC026356.1 by the online database. (F) Subcellular localization of AC026356.1 by FISH assay.
GO and KEGG analysis
To confirmed the potential function of AC026356.1 in LUAD. LncACTdb 3.0 database was used to obtain the positive genes with AC026356.1 in LUAD (Figure 4A, 4B). Moreover, we using the above genes conducted the GO and KEGG enrichment analysis. Results confirmed that AC026356.1 is major involved in the intermediate filament cytoskeleton, endonuclease activity, and nucleotide binding (Figure 4C). KEGG enrichment results showed that AC026356.1 is major involved in the hedgehog signaling (Figure 4D).
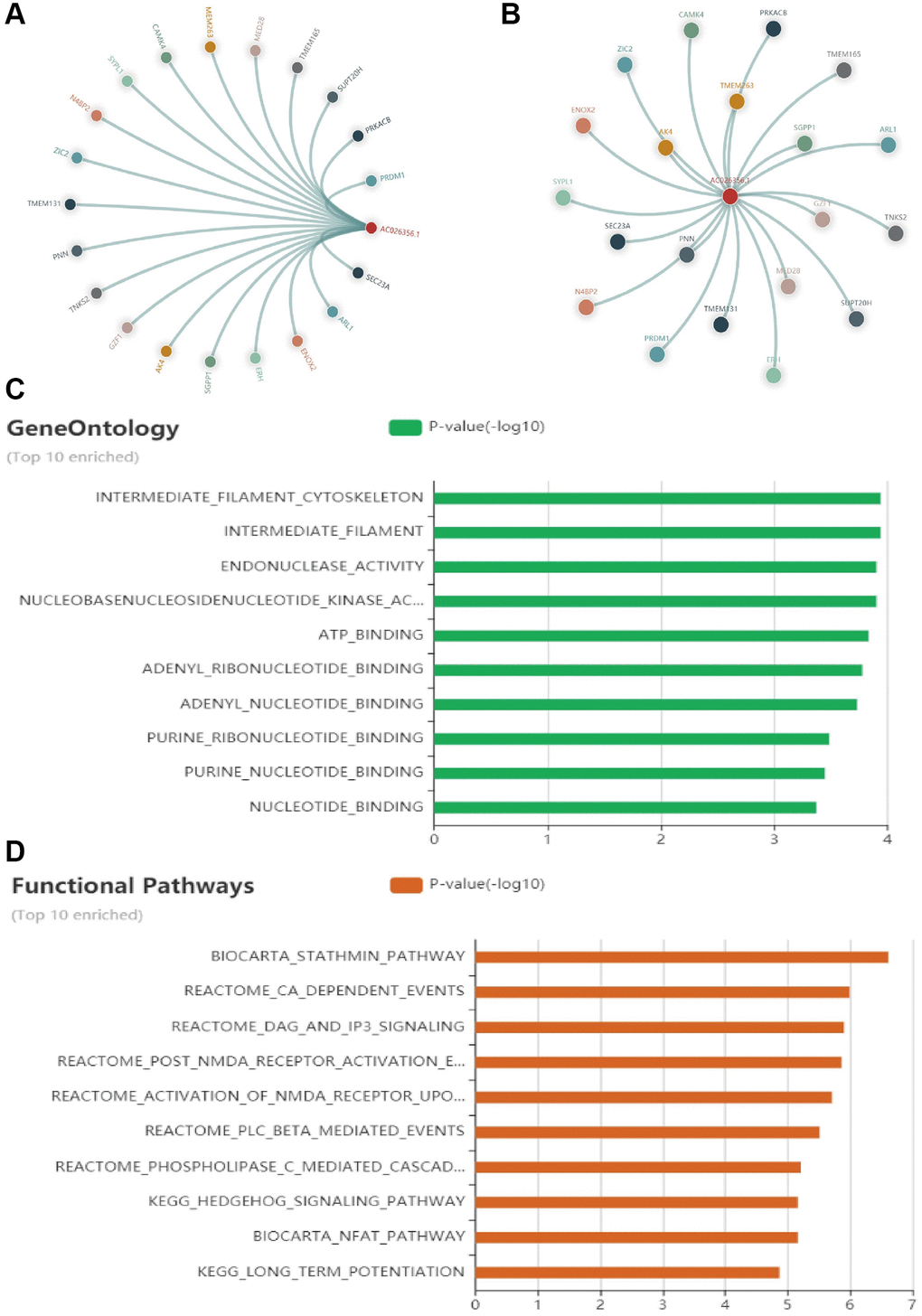
Figure 4. GO and KEGG enrichment analysis. (A, B) Positive genes with AC026356.1 in LUAD were examined by LncACTdb. (C, D) GO and KEGG enrichment analysis of AC026356.1 in LUAD examined by LncACTdb database.
AC026356.1 is required for malignant behaviors in LUAD cells
We confirmed that AC026356.1 was upregulated in LUAD cell lines than bronchial epithelial cell lines (Figure 5A). ASO using to inhibited AC026356.1 expression in lung cancer cells (Figure 5B). We uncover that reduced the expression of AC026356.1 was inhibited the cell growth and migration abilities of lung cancer cells (Figure 5C–5F).
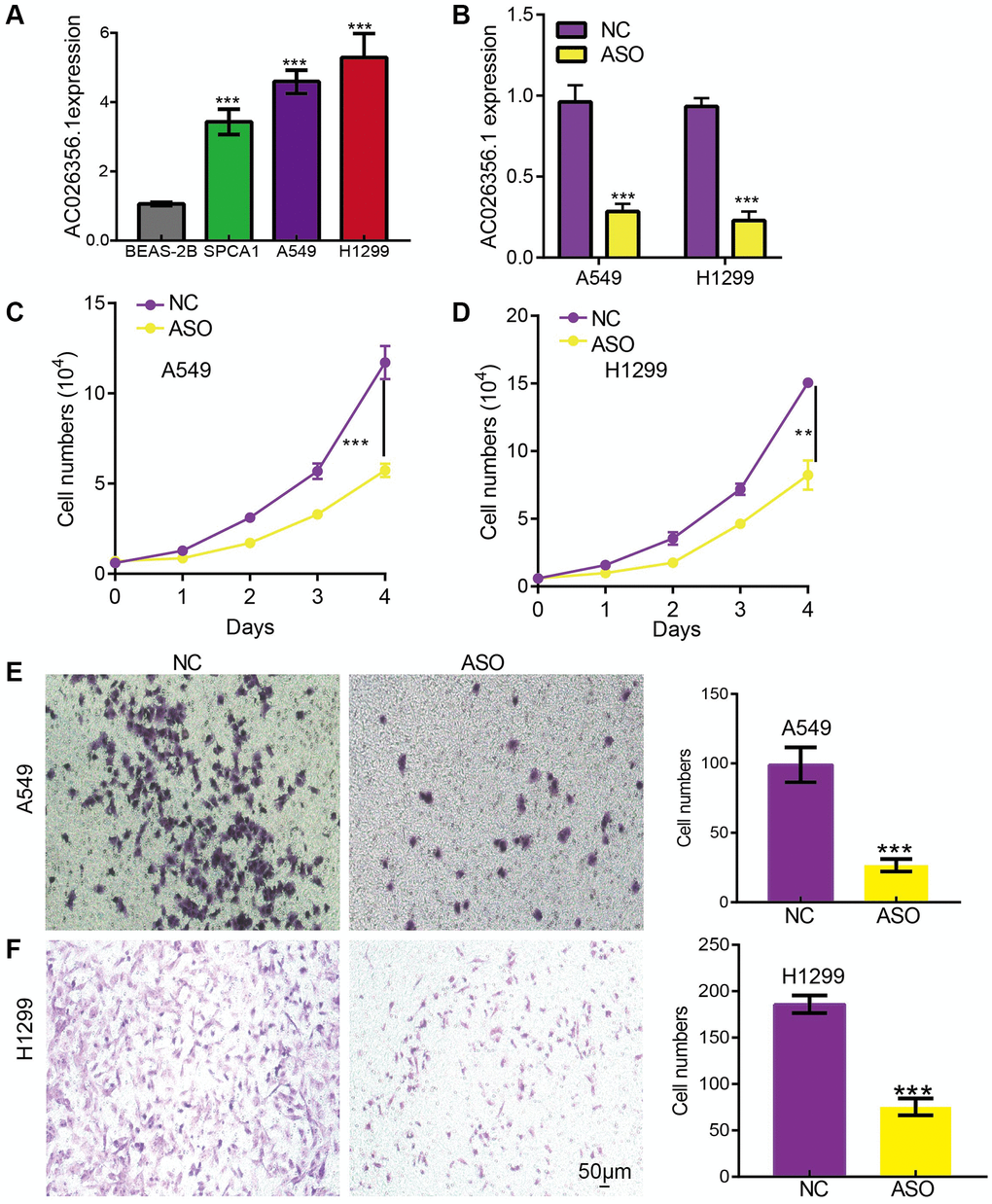
Figure 5. AC026356.1 promotes LUAD progression. (A) The expression of AC026356.1 in LUAD cell lines. (B) The knockdown efficiency was examined by qPCR assay. (C–F) Knockdown of AC026356.1 significantly inhibited LUAD cell growth and migration. **p < 0.01 and ***p < 0.001.
AC026356.1 is required for the self-renewal of cancer stem cell
Wnt signaling pathways play a core role in maintaining the stemness of cancer stem cells [14]. Given that GSEA enrichment results show that AC026356.1 was participated in the Wnt signaling pathway (Figure 6A). By analysis of the TCGA-LUAD data, we found that AC026356.1 expression was significantly positive with the stem cell-related gene, including CD133, CD44, SOX2, and OCT4 in LUAD (Figure 6B). we further investigated how AC026356.1 functions in lung cancer stemness. AC026356.1 knockdown repression significantly attenuated sphere formation of lung cancer cells (Figure 6C, 6D). Finally, we also showed that a AC026356.1 ceRNA regulated network in the progression of LUAD (Figure 6E, 6F). These data suggested that AC026356.1 modulate the stemness of LUAD cells.
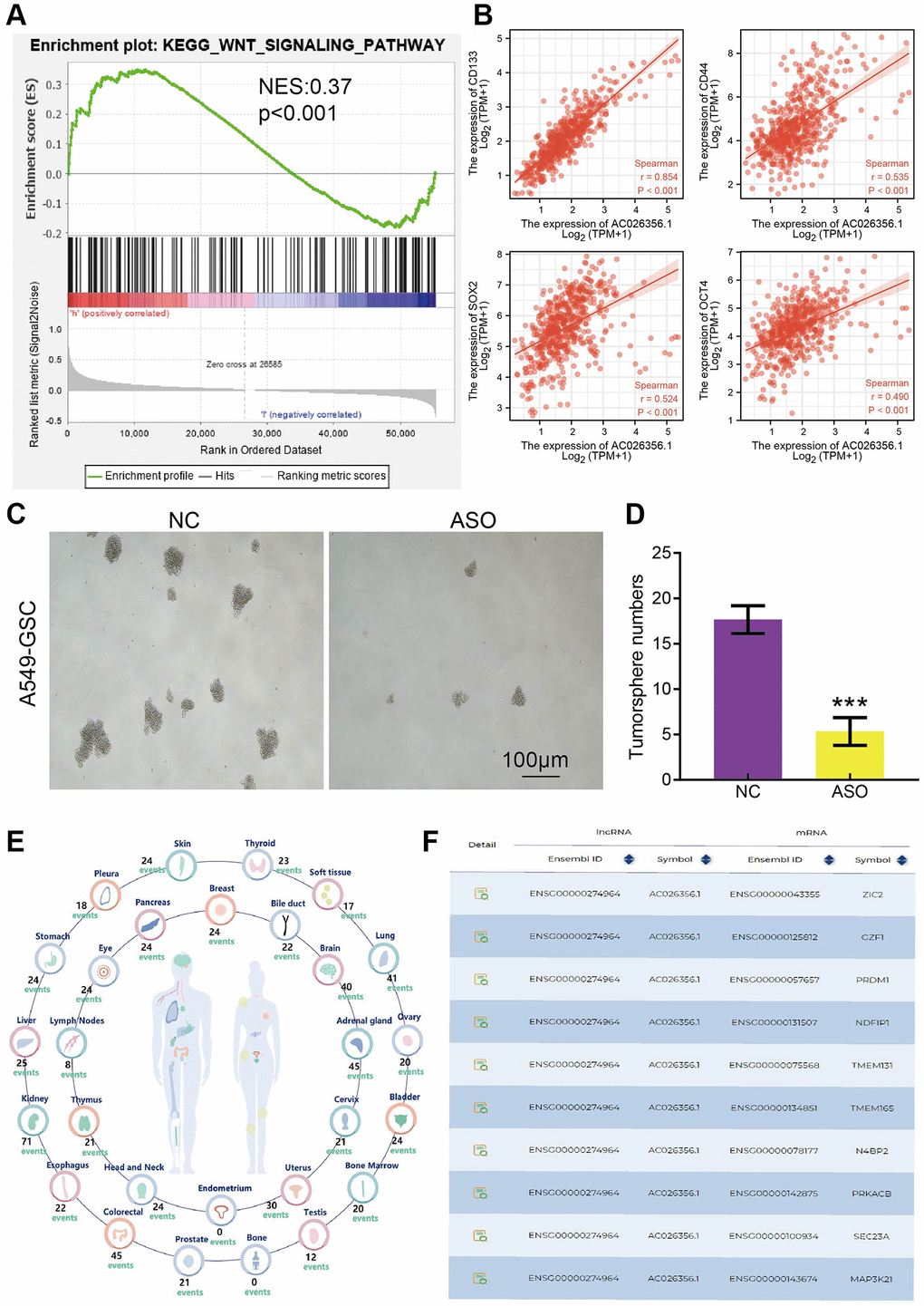
Figure 6. AC026356.1 is required for the self-renewal of cancer stem cell. (A) GSEA enrichment results show that AC026356.1 was involved in the Wnt signaling pathway. (B) Correlation between AC026356.1 and CD133, CD44, SOX2 and OCT4 in LUAD. (C, D) AC026356.1 knockdown repression significantly attenuated sphere formation of lung cancer cells. (E, F) The gene regulation by the lncRNA-miRNA-mRNA ceRNA network in the progression of LUAD. ***p < 0.001.
Correlation of AC026356.1 expression with LUAD immune cell infiltrates
The dynamic changes in the immune microenvironment play a core role in tumor progression [15]. We will further explore the correlations between AC026356.1 expression and the TME of LUAD. First, we showed that AC026356.1 expression was positively related to the LUAD immune scores, stromal scores, and ESTIMATE scores (Figure 7A). High expression of AC026356.1 was positively related to the infiltration levels of Th1 and Tem cells in LUAD (Figure 7B–7E). T-cell exhaustion is considered to be an important factor in the malignant progression of tumors [16]. More importantly, we uncover that AC026356.1 was negatively correlated with T cell exhaustion (Figure 7F). Finally, we found that in the AC026356.1 -high expression group, the infiltration levels of Macrophage, Th1 cells, Th17, Th2, Tem, T helper, macrophage, ADC, and NK cell were significantly increased than AC026356.1-low expression group (Figure 7G, 7H). We also confirmed that AC026356.1 expression was significantly positively linked to CD274, CTLA4, PDCD1, TIGIT, PDCD1LG2, HAVCR2, and SIGLEC15 in LUAD (Figure 7I).
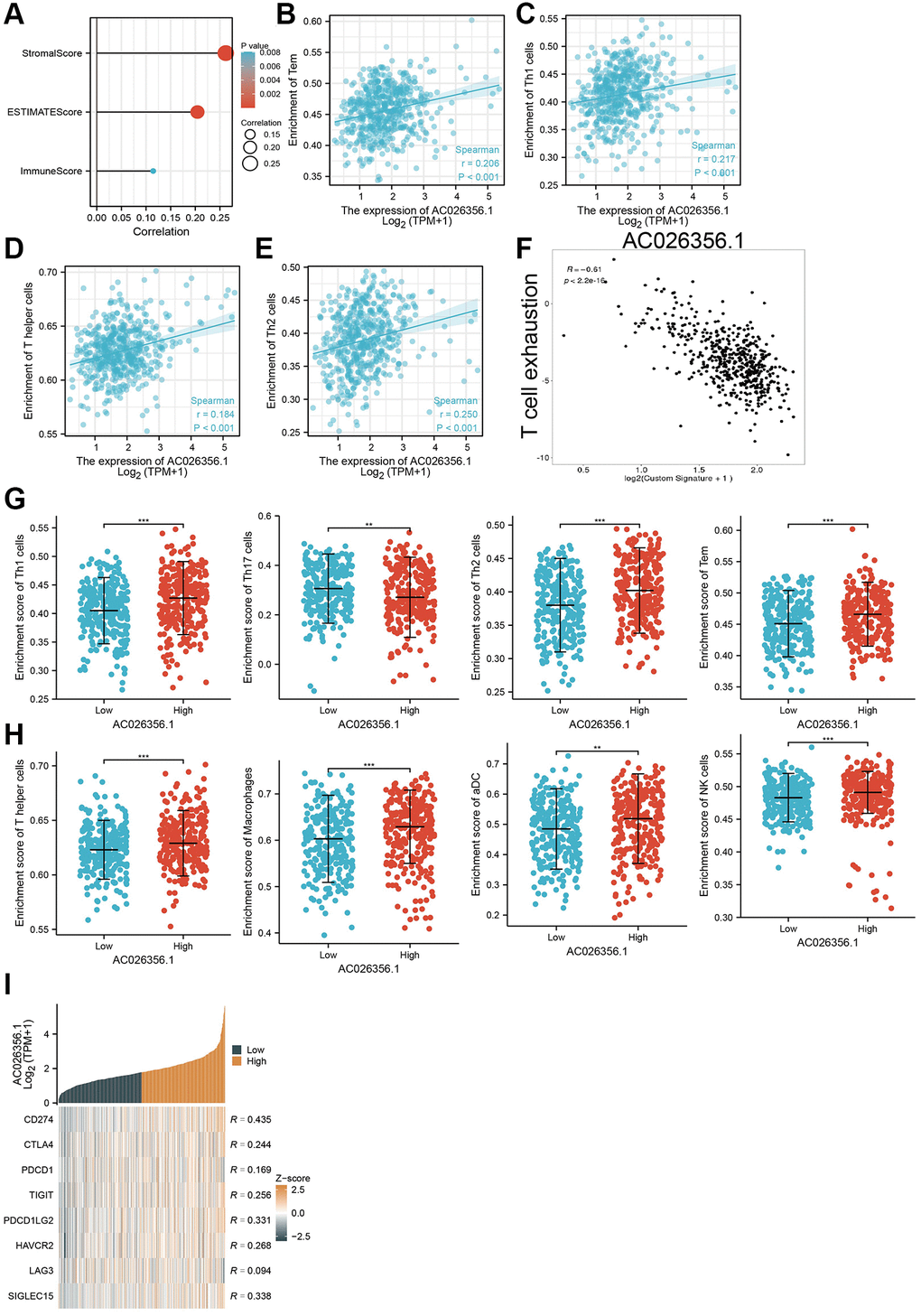
Figure 7. Correlation between AC026356.1 expression and immune infiltrates. (A) AC026356.1 expression was positively correlated with LUAD immune scores, stromal scores, and ESTIMATE scores. (B–E) Correlation between AC026356.1 expression and diverse immune infiltrates in LUAD. (F) Correlation between AC026356.1 expression and T cell exhaustion (G, H) The abundance of immune infiltrates of diverse immune cells based on the AC026356.1 high or low expression group. (I) Correlation between AC026356.1 expression and diverse immune checkpoint gene in LUAD. **p < 0.01 and ***p < 0.001.
Discussion
In this finding, we uncover that AC026356.1 expression was decreased in LUAD, compared to normal lung tissues. Meanwhile, low AC026356.1 expression is associated with the cancer stage and overall event in LUAD. Prognosis analysis indicated that LUAD patients with high AC026356.1 levels were related to adverse OS and DSS in LUAD. Our results demonstrated that AC026356.1 can be used as a prognostic biomarker for LUAD.
In this study, expression level and phenotypic analyses (including proliferation and sphere formation assays revealed that AC026356.1 formed a positive loop to regulate the stemness of lung cancer. Here, we unveiled METTL14/IGF2BP2-mediated m6A modification of AC026356.1, which lead to its high expression in lung cancer. Mechanistic disruption of the METTL14/IGF2BP2/AC026356.1 loop provided experimental evidence of how the positive loop was formed and exerted stemness upon m6A modification. In our study, we showed that AC026356.1 is upregulated in LUAD cells more than in the bronchial epithelial cell line. In vitro cell experiments indicated that depletion of AC026356.1 significantly inhibited LUAD cell growth and migration.
Recent studies have reported that the three m6A modifiers investigated in this study promote the propagation of various cancer cells including lung cancer cells. METTL14 is highly expressed in distinct tumor types and boosts stemness by enhancing SOX2 mRNA stability upon m6A modification [17]. IGF2BP2 interacts with and stabilizes FEN1 mRNA from decay in liver cancer [18]. Our data revealed that METTL14/IGF2BP2-mediated m6A modification of AC026356.1 RNA rescued them from decay. The existence of cancer stemness is the main cause of drug resistance, and distal metastasis- and CSC-targeted therapies hold promise in the clinical treatment of cancers. AC026356.1 is also essential for stemness-related malignancy in lung cancer. In vitro experiments indicated that AC026356.1 activation leads to resistance of NSCLC cells to erlotinib treatment.
The lung cancer microenvironment depends on the complex interactions between tumor cells and diverse immune cells. TME is thought to be a key factor contributing to tumor progression and high mortality in patients [19, 20]. By performing the correlation analysis, we showed that AC026356.1 expression is associated with immune scores, stromal scores, and ESTIMATE scores. High expression of AC026356.1 was positively related to the infiltration levels of Th1 and T cell exhaustion.
The present study provides insights to improve CSC-targeted therapy in cancer. Combination therapy of chemo drugs or targeting drugs with inhibitors or nucleic acid drugs targeting the IGF2BP2/AC026356.1 loop in vivo may further help in identifying novel therapeutic sensitizers for multiple regimens of cancer treatment.
Author Contributions
Zhen Zhang, Xiaoning Tan and Ruoxia Wu designed this work and performed related assays. Tianhao Deng and Huazhong Wang analysis of data. Xiulin Jiang, Puhua Zeng and Junqi Tang supervised and wrote the manuscript. All authors have read and approved the final version of the manuscript.
Acknowledgments
The authors would like to thank the TCGA and GTEx databases for providing the data.
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
Ethical Statement
This study was approved by the Ethics Committee of Affiliated Hospital of Hunan Academy of Traditional Chinese Medicine.
Funding
This work was supported by Yunnan Province Specialized Training Grant for High-Level Healthcare Professionals (D-201614).
Editorial Note
This corresponding author has a verified history of publications using a personal email address for correspondence.
References
- 1. Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. CA Cancer J Clin. 2021; 71:7–33. https://doi.org/10.3322/caac.21654 [PubMed]
- 2. Gray SG, Mutti L. Immunotherapy for mesothelioma: a critical review of current clinical trials and future perspectives. Transl Lung Cancer Res. 2020; 9:S100–19. https://doi.org/10.21037/tlcr.2019.11.23 [PubMed]
- 3. Guazzelli A, Meysami P, Bakker E, Demonacos C, Giordano A, Krstic-Demonacos M, Mutti L. BAP1 Status Determines the Sensitivity of Malignant Mesothelioma Cells to Gemcitabine Treatment. Int J Mol Sci. 2019; 20:429. https://doi.org/10.3390/ijms20020429 [PubMed]
- 4. Yang J, Liu F, Wang Y, Qu L, Lin A. LncRNAs in tumor metabolic reprogramming and immune microenvironment remodeling. Cancer Lett. 2022; 543:215798. https://doi.org/10.1016/j.canlet.2022.215798 [PubMed]
- 5. Ricciardi S, Cardillo G, Zirafa CC, Carleo F, Facciolo F, Fontanini G, Mutti L, Melfi F. Surgery for malignant pleural mesothelioma: an international guidelines review. J Thorac Dis. 2018; 10:S285–92. https://doi.org/10.21037/jtd.2017.10.16 [PubMed]
- 6. He Y, Jiang X, Duan L, Xiong Q, Yuan Y, Liu P, Jiang L, Shen Q, Zhao S, Yang C, Chen Y. LncRNA PKMYT1AR promotes cancer stem cell maintenance in non-small cell lung cancer via activating Wnt signaling pathway. Mol Cancer. 2021; 20:156. https://doi.org/10.1186/s12943-021-01469-6 [PubMed]
- 7. Liang R, Li X, Li W, Zhu X, Li C. DNA methylation in lung cancer patients: Opening a "window of life" under precision medicine. Biomed Pharmacother. 2021; 144:112202. https://doi.org/10.1016/j.biopha.2021.112202 [PubMed]
- 8. Peiris-Pagès M, Martinez-Outschoorn UE, Pestell RG, Sotgia F, Lisanti MP. Cancer stem cell metabolism. Breast Cancer Res. 2016; 18:55. https://doi.org/10.1186/s13058-016-0712-6 [PubMed]
- 9. Najafi M, Mortezaee K, Majidpoor J. Cancer stem cell (CSC) resistance drivers. Life Sci. 2019; 234:116781. https://doi.org/10.1016/j.lfs.2019.116781 [PubMed]
- 10. Huang H, Weng H, Chen J. m6A Modification in Coding and Non-coding RNAs: Roles and Therapeutic Implications in Cancer. Cancer Cell. 2020; 37:270–88. https://doi.org/10.1016/j.ccell.2020.02.004 [PubMed]
- 11. Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, Zhao BS, Mesquita A, Liu C, Yuan CL, Hu YC, Hüttelmaier S, Skibbe JR, et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018; 20:285–95. https://doi.org/10.1038/s41556-018-0045-z [PubMed]
- 12. Wang P, Guo Q, Qi Y, Hao Y, Gao Y, Zhi H, Zhang Y, Sun Y, Zhang Y, Xin M, Zhang Y, Ning S, Li X. LncACTdb 3.0: an updated database of experimentally supported ceRNA interactions and personalized networks contributing to precision medicine. Nucleic Acids Res. 2022; 50:D183–9. https://doi.org/10.1093/nar/gkab1092 [PubMed]
- 13. Jin D, Guo J, Wu Y, Du J, Yang L, Wang X, Di W, Hu B, An J, Kong L, Pan L, Su G. m6A mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. J Hematol Oncol. 2019; 12:135. https://doi.org/10.1186/s13045-019-0830-6 [PubMed]. Retraction in: J Hematol Oncol. 2023; 16:14. [PubMed]
- 14. Duchartre Y, Kim YM, Kahn M. The Wnt signaling pathway in cancer. Crit Rev Oncol Hematol. 2016; 99:141–9. https://doi.org/10.1016/j.critrevonc.2015.12.005 [PubMed]
- 15. Qiao M, Jiang T, Liu X, Mao S, Zhou F, Li X, Zhao C, Chen X, Su C, Ren S, Zhou C. Immune Checkpoint Inhibitors in EGFR-Mutated NSCLC: Dusk or Dawn? J Thorac Oncol. 2021; 16:1267–88. https://doi.org/10.1016/j.jtho.2021.04.003 [PubMed]
- 16. Wherry EJ. T cell exhaustion. Nat Immunol. 2011; 12:492–9. https://doi.org/10.1038/ni.2035 [PubMed]
- 17. Yang X, Zhang S, He C, Xue P, Zhang L, He Z, Zang L, Feng B, Sun J, Zheng M. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol Cancer. 2020; 19:46. https://doi.org/10.1186/s12943-020-1146-4 [PubMed]
- 18. Pu J, Wang J, Qin Z, Wang A, Zhang Y, Wu X, Wu Y, Li W, Xu Z, Lu Y, Tang Q, Wei H. IGF2BP2 Promotes Liver Cancer Growth Through an m6A-FEN1-Dependent Mechanism. Front Oncol. 2020; 10:578816. https://doi.org/10.3389/fonc.2020.578816 [PubMed]
- 19. Thomas PD, Kahn M. Kat3 coactivators in somatic stem cells and cancer stem cells: biological roles, evolution, and pharmacologic manipulation. Cell Biol Toxicol. 2016; 32:61–81. https://doi.org/10.1007/s10565-016-9318-0 [PubMed]
- 20. Kothari AN, Mi Z, Zapf M, Kuo PC. Novel clinical therapeutics targeting the epithelial to mesenchymal transition. Clin Transl Med. 2014; 3:35. https://doi.org/10.1186/s40169-014-0035-0 [PubMed]