Introduction
Lung cancer is the most lethal tumor of respiratory system [1]. Lung adenocarcinomas are found in nearly 40% of lung cancers, and squamous cell carcinomas and large cell carcinomas are the other two subtypes of NSCLC [2]. For patients with lung adenocarcinoma (LUAD), the overall survival times is less than 20% [2]. It is very important to study the initiation and metastasis of lung cancer and identify new biomarkers for the prevention and diagnosis of lung cancer.
DTYMK, a nuclear deoxythymidylate kinase, plays an important role in catalyzes the process of deoxy-TMP phosphorylation. Relevant studies have reported that DTYMK exerts a pivotal effect on multiple physiological processes including genome integrity and neuronal survival [3]. It has been reported that DTYMK was dramatically upregulated in HCC and correlated with worse overall survival [4]. However, the potential biological and clinical value of DTYMK in lung cancer remains unclear.
In this finding, we comprehensively analysis the expression patterns, clinical relevance and prognostic value of DTYMK in lung cancer. Finally, the biological event of DTYMK in lung cancer progression was examined by loss of function. Our findings may provide insight into the mechanism and role of DTYMK in LUAD.
Materials and Methods
Data processing
The normalized gene expression matrix and clinical information data in various tumor and normal tissue samples was downloaded from TCGA database and GenotypeTissue Expression (GTEx) databases.
Gene enrichment analysis based on DTYMK co-expressed genes
We examined the co-expression genes of DTYMK by using LinkedOmics (http://www.linkedomics.org/login.php) [5]. We also performed the GO and KEGG enrichment analysis of DTYMK in LUAD by using the R package “clusterProfiler”.
Diagnostic and prognosis values
In this manuscript, we mainly used the ROC curve and Kaplan-Meier Plotter to examine the potential diagnostic and prognosis values of DTYMK in LUAD [6].
Immune infiltration analysis of DTYMK in LUAD
The TIMER database is a bioinformatics web tool for comprehensively analyzing tumor-infiltrating immune cells (TIICs) [7]. The correlation between DTYMK and various human immune cells in LUAD was analyzed in TIMER.
Loss of function assay
We conducted the CCK8 assay and transwell assay to examine the potential biological function of DTYMK on the cell behavior of LUAD. Above assay mainly refer to previous and published literature [8].
Statistical analyses
The statistical for Figure analyses were performed using R (V 3.6.3), and ROC curves were used to detect DTYMK cutoff values using pROC packages. P < 0.05 (*), P < 0.01 (**) and P < 0.001 (***), were significant.
Results
DTYMK was over-expression in pan-cancer
We used TCGA and GTEx datasets to examine the levels of DTYMK in various cancer types. Results confirmed that DTYMK was up-regulated in many human cancer (Figure 1A, 1B). We also found that DTYMK expression was significantly increased compared with normal tissues in both unpaired and paired expression (Figure 2A, 2B). In addition, we showed that DTYMK protein level was elevated in LUAD than in normal tissue (Figure 2C, 2D).
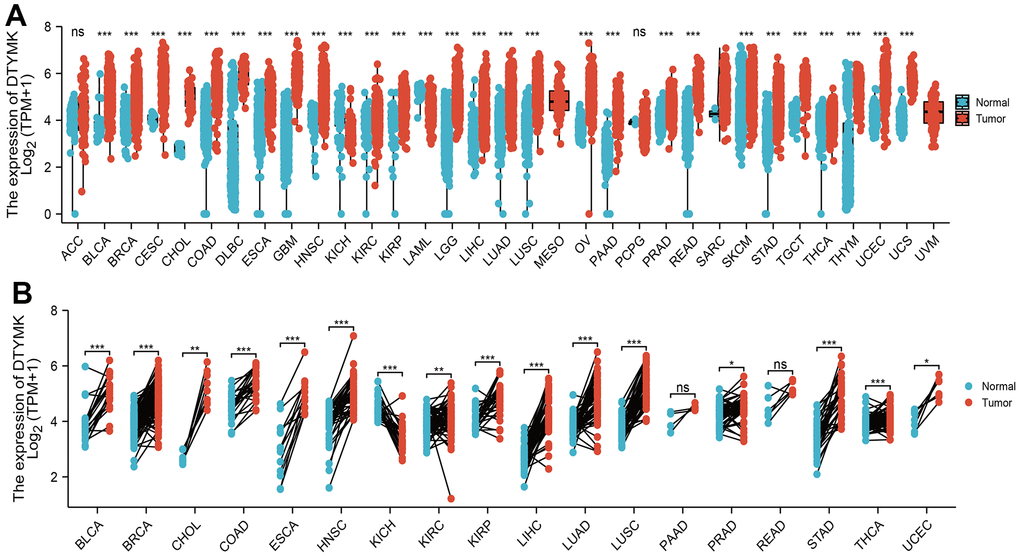
Figure 1. Expression pattern of DTYMK from the perspective of pan-cancer. (A) DTYMK was highly expressed in 24 of the 33 cancers compared with normal tissue. (B) Relative DTYMK expression in paired lung cancer and noncancerous tissues examined by TCGA databases. ACC: Adrenocortical carcinoma, BLCA: Bladder Urothelial Carcinoma, BRCA: Breast invasive carcinoma, CESC: Cervical squamous cell carcinoma and endocervical adenocarcinoma, CHOL: Cholangiocarcinoma, COAD: Colon adenocarcinoma, DLBC: Lymphoid Neoplasm Diffuse Large B-cell Lymphoma, ESCA: Esophageal carcinoma, GBM: Glioblastoma multiforme, HNSC: Head and Neck squamous cell carcinoma, KICH: Kidney: Chromophobe, KIRC: Kidney renal clear cell carcinoma, KIRP: Kidney renal papillary cell carcinoma, LAML: Acute Myeloid Leukemia, LUAD: Brain Lower Grade Glioma, LIHC: Liver hepatocellular carcinoma, LUAD: Lung adenocarcinoma, LUSC: Lung squamous cell carcinoma, MESO: Mesothelioma, OV: Ovarian serous cystadenocarcinoma, PAAD: Pancreatic adenocarcinoma, PCPG: Pheochromocytoma and Paraganglioma, PRAD: Prostate adenocarcinoma, READ: Rectum adenocarcinoma, SARC: Sarcoma, SKCM: Skin Cutaneous Melanoma, STAD: Stomach adenocarcinoma, TGCT: Testicular Germ Cell Tumors, THCA: Thyroid carcinoma, THYM: Thymoma, UCEC: Uterine Corpus Endometrial Carcinoma, UCS: Uterine Carcinosarcoma, UVM: Uveal Melanoma. NS: P >0.05,*P < 0.05, **P < 0.01, ***P < 0.001.
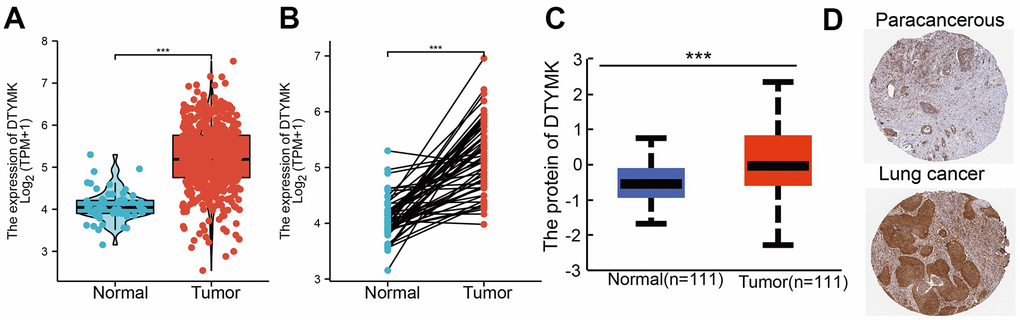
Figure 2. DTYMK RNA and protein expression in LUAD. (A) DTYMK mRNA expression levels in 535 lung cancer patients and matched adjacent normal samples. (B) Relative DTYMK expression in 54 paired lung cancer and noncancerous tissues. (C) DTYMK protein expression level based on CPTAC. (D) DTYMK protein levels based on Human Protein Atlas. ***P < 0.001.
PPI network and functional annotation
As shown in Figure 6A, STRING database utilized to construct the PPI network. Furthermore, we also obtained the top co-expressed genes (Figure 6B–6D). Functional analysis showed that DTYMK mainly related to Oxidative phosphorylation, Cell cycle, Proteasome, 53 signaling pathway, RNA degradation, and RNA polymerase (Figure 6E). GSEA result show that DTYMK mainly involved in cell cycle, Oxidative phosphorylation, Focal adhesion, and chemokine signaling pathway (Figure 7A, 7B).
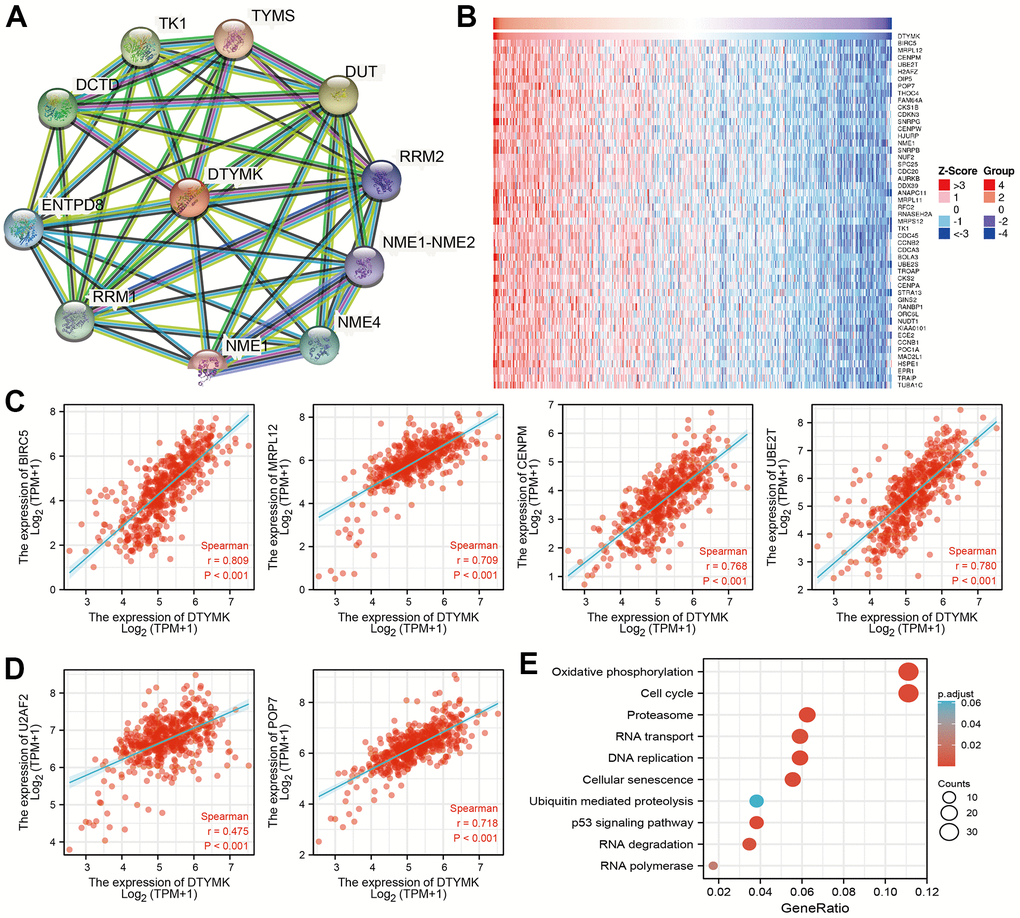
Figure 6. PPI network and functional enrichment analysis. (A) The protein-protein network of DTYMK examine by STRING database. (B–D) The correlation analysis of DTYMK expression and its top 100 co-expressed gene network. (E) Functional enrichment analysis of co-expressed genes.
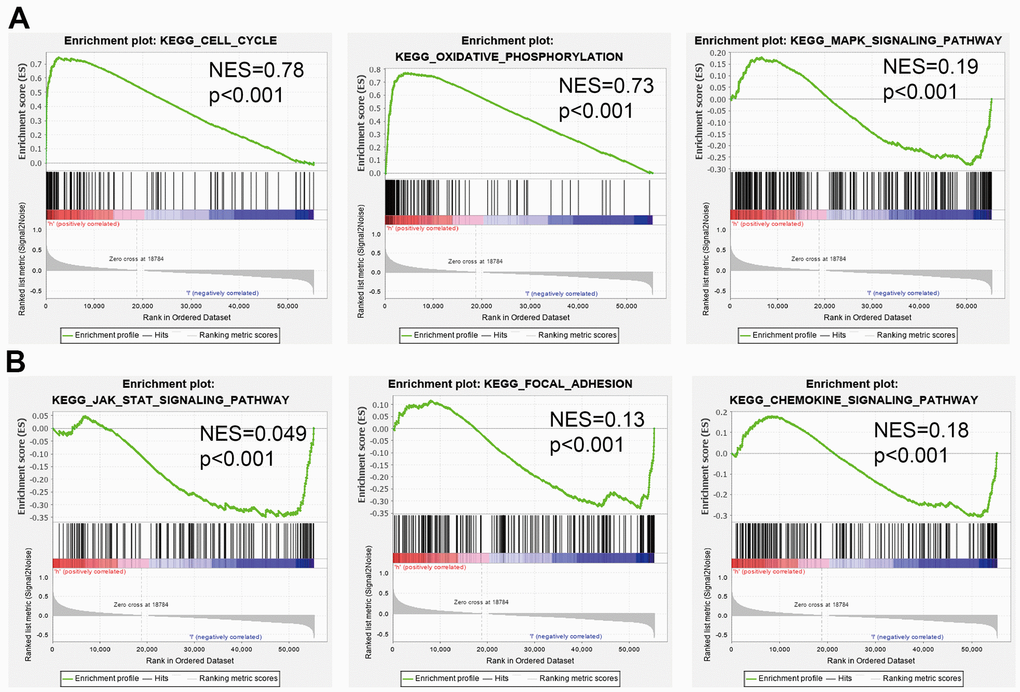
Figure 7. Identification of DTYMK related signaling pathways in lung adenocarcinoma. (A, B) Identification of DTYMK related signaling pathways by GSEA software.
Immune infiltration analysis of DTYMK
Firstly, the TIMER database was employed to evaluate the relationship between DTYMK levels and the level of TIICs infiltration in TCGA-LUAD. Different infiltration levels of TIICs appeared to be related to the alteration of DTYMK, including various immune cells (Figure 8A). Moreover, we show that DTYMK was significantly correlated with the level of diverse human immune cells (Figure 8B). Finally, we determined that elevated DTYMK expression was positive related to the abundance of 22 human immune cells (Figure 8C–8E).
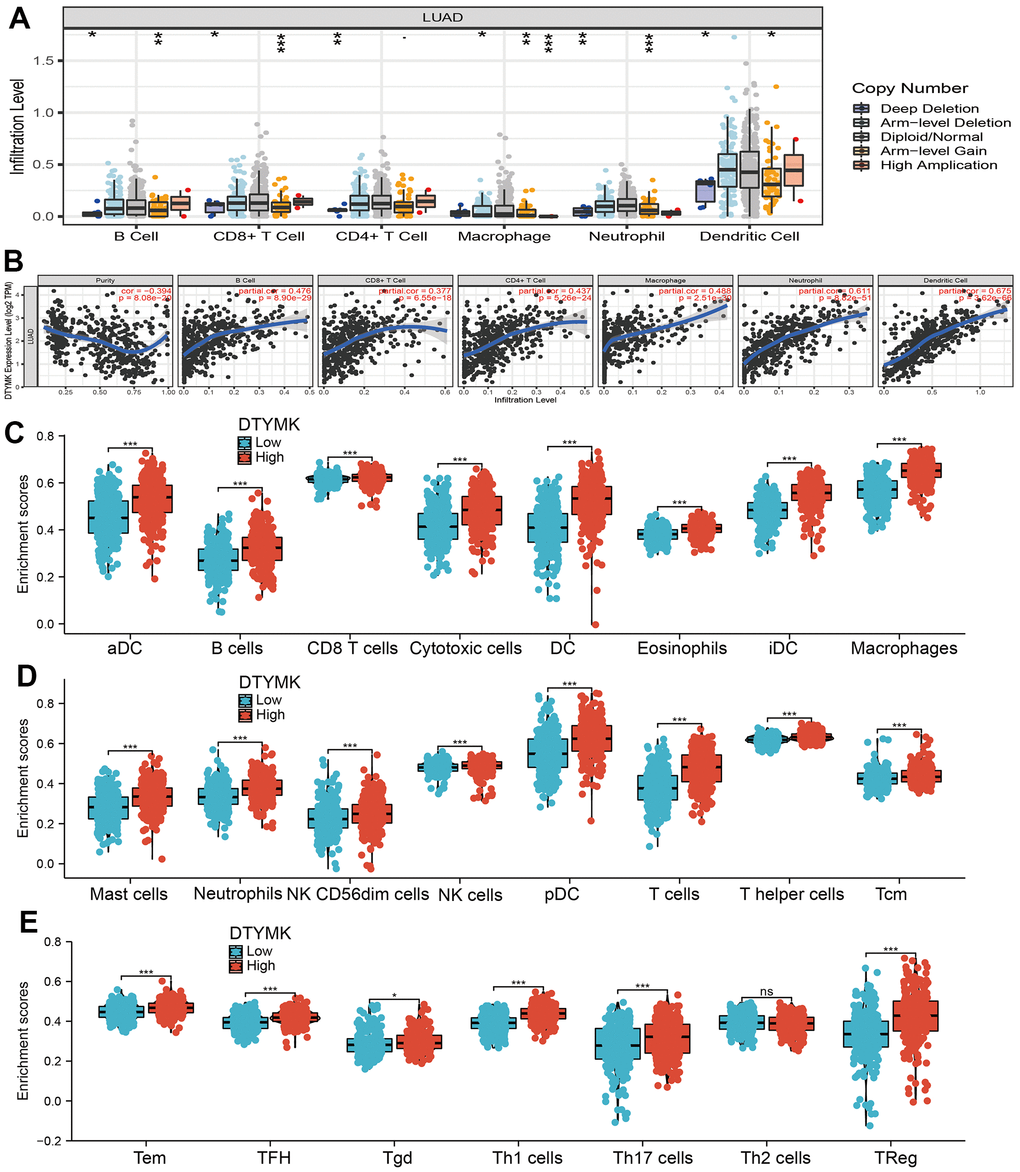
Figure 8. Correlation analysis of DTYMK expression and infiltration levels of immune cells in LUAD. (A) The correlation between DTYMK expression and somatic copy number alterations examine by TIMER. (B) The correlation between DTYMK expression and the infiltration levels of B cells, CD4+ T cells, CD8+ T cells, dendritic cells, Macrophages and Neutrophils. (C–E) Box plots of the correlations between DTYMK or molecular model expression and infiltration levels of immune cells. NS: P >0.05,*P < 0.05, **P < 0.01, ***P < 0.001.
Knockdown of DTYMK inhibited LUAD progression
We showed that DTYMK was up-regulated in A549 and H1975 cells (Figure 9A, 9B). Loss of function assays suggested that knockdown of DTYMK inhibited the cell growth and migration abilities of LUAD cells (Figure 9C–9E). Furthermore, DTYMK knock-down led to increased G0/G1 phase arrested cells (Figure 9F).
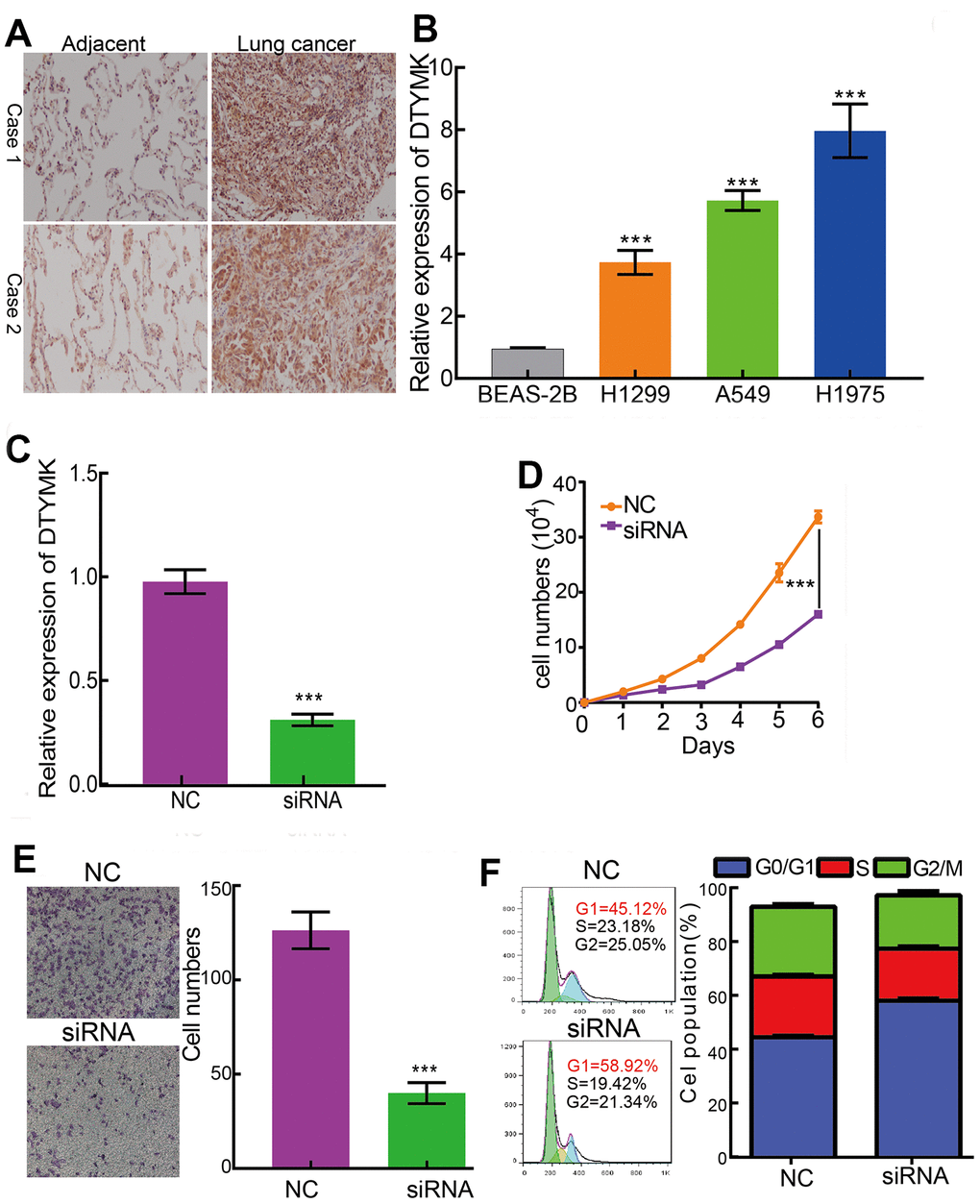
Figure 9. DTYMK regulates LUAD cell proliferation, migration and cell cycle. (A) IHC assay detect the protein of DTYMK in lung adenocarcinoma cancerous. (B) qPCR assay examine the expression level of DTYMK in lung adenocarcinoma cancerous cell lines, including A549, H1299, and H1975, compared to normal human bronchial epithelial cell line: BEAS-2B. (C) Establishment of DTYMK knockdown cell lines in H1975 verified by Real-time RT-PCR (D, E) Knockdown of DTYMK significantly inhibits cell proliferation and migration in H1975 cells, as measured by CCK8 and transwell assays. (F) Depletion of DTYMK increased G0/G1 phase arrested cells. NC=negative control, siRNA= DTYMK siRNA ***p < 0.001.
Discussion
It has been confirmed that DTYMK mainly catalyzes the last reaction of the deoxyribonucleoside triphosphate (dTTP) production pathways [9]. According to previous studies, we showed that DTYMK was significantly different in LUAD and normal tissues and associated with OS in LUAD patients.
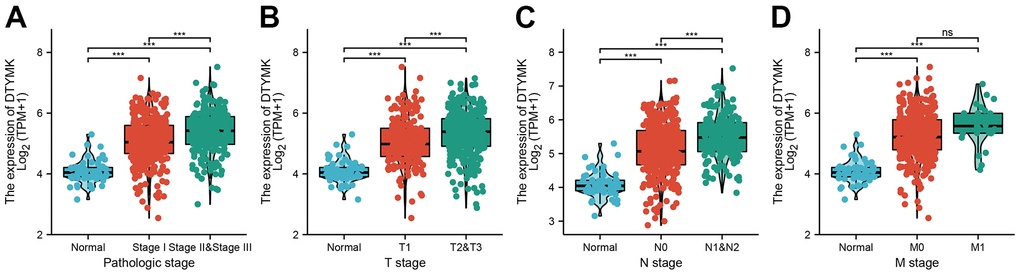
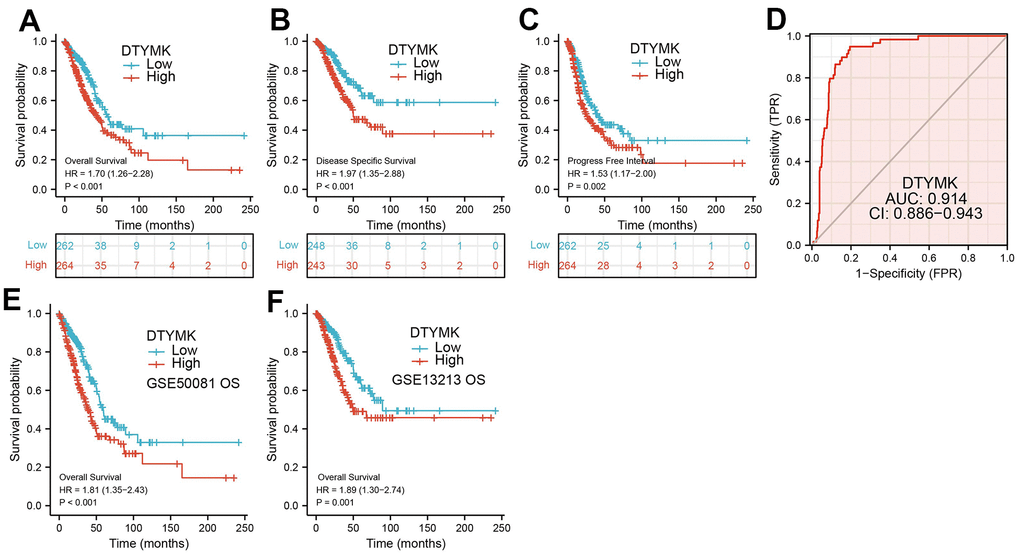
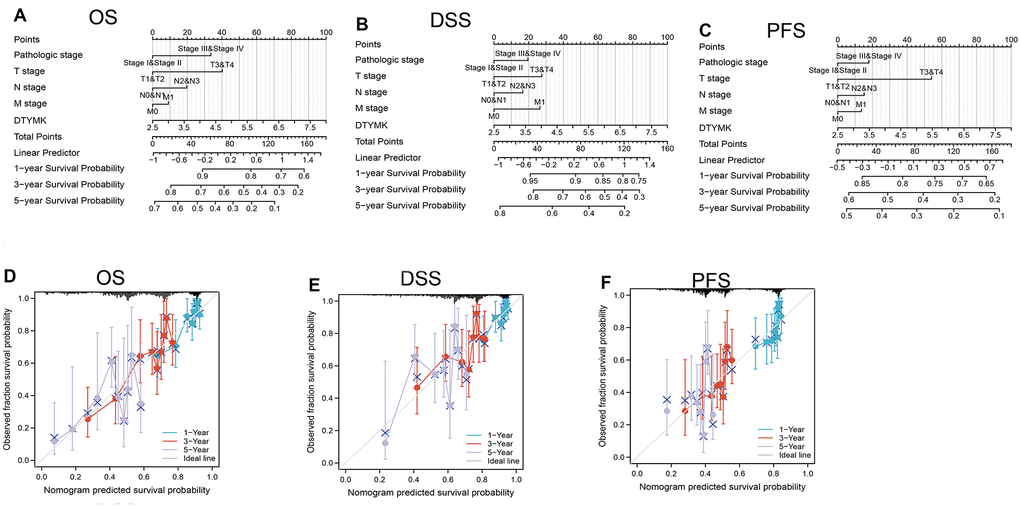
In this finding, we evaluated the expression of DTYMK in LUAD dependent databases such as TCGA, GTEx, and UALCAN. We found that DTYMK was up-regulated in LUAD. Moreover, DTYMK expression was increased in the different cancer stages of LUAD, compared with the normal control group. Next, to examine whether DTYMK can be used as a prognostic biomarker, we analyzed the relationship between DTYMK in LUAD and OS, DSS, and PFI and plotted the survival curve using Kaplan-Meier plotter databases. Notably, increased DTYMK expression was associated with worse OS, DSS, and PFS. Above results showed that up-regulation of DTYMK was associated with a worse prognosis in LUAD patients.
Previous studies reported that DTYMK was significant for neuronal survival [3]. In this finding, we show that DTYMK are involved in Oxidative phosphorylation, Proteasome, 53 signaling pathway, RNA degradation, and RNA polymerase.
Our study showed that DTYMK is closely correlated with immune cells, suggesting that DTYMK plays a critical role in lung cancer immunity. Immune checkpoints are pairs of receptor-ligand molecules that interact to suppress immune responses, and tumors can protect themselves by using immune checkpoints to evade immune clearance [10]. Our study also found that DTYMK is associated with immune in human lung cancer types. In conclusion, DTYMK may affect the prognosis of various cancers and is related to immune infiltration.
We have analyzed the role of DTYMK in LUAD and the potential clinical value of DTYMK in lung cancer. However, the current study still had some limitations. First, there were no in vivo experiments to verify the role of DTYMK in LUAD. Overall, our results confirmed that DTYMK may be a potential novel prognostic biomarker for LUAD, which may improve our understanding of the translational use of DTYMK in LUAD prognosis and therapy.
Author Contributions
Xi Chen, Yixiao Yuan and Fan Zhou designed this work, performed related assay. Xiaobin Huang and Jun Pu analyzed data. Xiaoqun Niu, Xiulin Jiang and Xiaoyan Ding supervised and wrote the manuscript. All authors have read and approved the final version of the manuscript.
Acknowledgments
We would like to thank the Core Technology Facility, Kunming Institute of Zoology, Chinese Academy of Sciences for providing us with bioinformatics analysis, and we are grateful to Guolan Ma for her technical support.
Conflicts of Interest
The authors declare no conflicts of interest.
Ethical Statement and Consent
The study of patients’ tissue samples was reviewed and approved by the Medical Ethics and Human Clinical Trial Committee of the Second Affiliated Hospital of Kunming Medical University. Written informed consent was provided by all patients before sample collection.
Funding
This work was supported by the National Nature Science Foundation of China (82160512, 30960398, 81460174, 81360126), Yunnan Applied Basic Research Projects (2017FE467 and 2018FE001), and the Applied Basic Research Project of Yunnan Provincial Science and Technology Department, Kunming Medical University (No.2020001AY070001-117) and the Open Project of The First People's Hospital of Yunnan Province Clinical Medicine Center (2021LCZXXF-XZ03).
Editorial Note
This corresponding author has a verified history of publications using a personal email address for correspondence
References
- 1. de Sousa VM, Carvalho L. Heterogeneity in Lung Cancer. Pathobiology. 2018; 85:96–107. https://doi.org/10.1159/000487440 [PubMed]
- 2. Jonna S, Subramaniam DS. Molecular diagnostics and targeted therapies in non-small cell lung cancer (NSCLC): an update. Discov Med. 2019; 27:167–70. [PubMed]
- 3. Vanoevelen JM, Bierau J, Grashorn JC, Lambrichs E, Kamsteeg EJ, Bok LA, Wevers RA, van der Knaap MS, Bugiani M, Frisk JH, Colnaghi R, O’Driscoll M, Hellebrekers DM, et al. DTYMK is essential for genome integrity and neuronal survival. Acta Neuropathol. 2022; 143:245–62. https://doi.org/10.1007/s00401-021-02394-0 [PubMed]
- 4. Zhou T, Qin R, Shi S, Zhang H, Niu C, Ju G, Miao S. DTYMK promote hepatocellular carcinoma proliferation by regulating cell cycle. Cell Cycle. 2021; 20:1681–91. https://doi.org/10.1080/15384101.2021.1958502 [PubMed]
- 5. Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2018; 46:D956–63. https://doi.org/10.1093/nar/gkx1090 [PubMed]
- 6. Lánczky A, Győrffy B. Web-Based Survival Analysis Tool Tailored for Medical Research (KMplot): Development and Implementation. J Med Internet Res. 2021; 23:e27633. https://doi.org/10.2196/27633 [PubMed]
- 7. Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, Li B, Liu XS. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017; 77:e108–10. https://doi.org/10.1158/0008-5472.CAN-17-0307 [PubMed]
- 8. Hua Q, Jin M, Mi B, Xu F, Li T, Zhao L, Liu J, Huang G. LINC01123, a c-Myc-activated long non-coding RNA, promotes proliferation and aerobic glycolysis of non-small cell lung cancer through miR-199a-5p/c-Myc axis. J Hematol Oncol. 2019; 12:91. https://doi.org/10.1186/s13045-019-0773-y [PubMed]
- 9. Caspi R, Billington R, Ferrer L, Foerster H, Fulcher CA, Keseler IM, Kothari A, Krummenacker M, Latendresse M, Mueller LA, Ong Q, Paley S, Subhraveti P, et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 2016; 44:D471–80. https://doi.org/10.1093/nar/gkv1164 [PubMed]
- 10. Mayer-Barber KD, Barber DL. Innate and Adaptive Cellular Immune Responses to Mycobacterium tuberculosis Infection. Cold Spring Harb Perspect Med. 2015; 5:a018424. https://doi.org/10.1101/cshperspect.a018424 [PubMed]