Introduction
Bladder cancer (BC), a forth common cancer in men, counts on high incidence and mortality rate, estimated 549,000 new cases and 200,000 deaths in 2018, globally [1, 2]. Under the disease management of intravesical BCG instillations for NMIBC and radical cystectomy or chemotherapy for MIBC, or further several novel therapies such as immunotherapy, oncolytic viruses, it has been reported that the survival rate of bladder cancer within 5 years at all stages is no more than 20% [3]. Under the current diagnosis and treatment of bladder cancer, it is urgent to identify more reliable diagnosis and prognostic indicators.
Recent years, we’ve gradually come to know that the tumor microenvironment (TME) plays an essential role in tumor differentiation, tumor epigenetics, immune evasion, and even treatment resistance [4]. The state of host innate immune system and the proportion of local infiltration of different types of immune cells are critical factors of the TME [5]. Notably, the tumor immune micro-environment is similar with the immune infiltration of chronic inflammation, containing multiple different types of immune cells [6]. Immune infiltration in TME have been proved to have significant influence on both tumor-promoting and suppressing activities. Other studies also suggest that the density of immune cells is associated with the immune evasion and treatment resistance of breast cancer [7]. Among of these immune cells, lymphocytes are the main type of inflammatory immune cells in TME [8]. Besides, the infiltration of CD8+ T and CD4+ T cells is also reported with a significant influence on the prognosis of bladder carcinoma [9, 10]. Specifically, tumor infiltrating CD8+ T cells have anti-tumor function and show a positive effect on prognosis of many tumors [11–13]. Also, tumor associated macrophages play an essential role in bladder cancer, especially M2 macrophages [14]. As for dendritic cells (DC) and other antigen-presenting cells (APC), they play a significant role in the biological process of tumor antigen presentation [15, 16].
Long non-coding RNA (LncRNA) is a class of non-coding RNAs with transcripts longer than 200 nt, which do not translate proteins but regulate gene expression by multiple mechanisms in the form of RNA [17]. Meanwhile, lncRNAs play significant role in the progression and prognosis in human diseases, especially in cancer [18]. For example, in a recent meta-analysis, Quan, et al. reported that lncRNAs were associated with the prognosis of patients with bladder cancer and could perform as an effective prediction factor for the overall survival of BC patients [19]. Among these lncRNAs, UCA1 was considered to serve as an efficient biomarker in the diagnosis of bladder cancer, while the abnormal expression of HOTAIR and GAS5 was associated with a poor prognosis including elapse-free survival (RFS), disease-free survival (DFS), r and disease-specific survival (DSS) [19]. Furthermore, lncRNAs were reported to regulate the immune infiltration directly or indirectly [20]. For instance, it has been reported LNC-INSR could enhance Treg cells differentiation and promote immunosuppression in childhood acute lymphoblastic leukemia [21]. Recent studies have also shown that lncRNAs have a high frequency and cell-type-specific presence in different type of immune cells, and the expression pattern of key lncRNAs is also determined to be related to immune infiltration in TME [22].
Therefore, we aimed to screen immune-infiltration-related lncRNAs in BC patients. In this research, we comprehensively assess the immune infiltration of BC patients by two methods including single sample gene set enrichment analysis (ssGSEA) and ESTIMATE algorithm. Then identified differential immune subtype BC patients and discovered differentially expressed immune infiltration related lncRNAs. Finally, we developed a 12 differentially expressed immune-infiltrating related lncRNAs signature and demonstrated the correlation between risk score calculated by the signature and tumor microenvironment in bladder cancer.
Results
Identification of bladder cancer immune subgroups
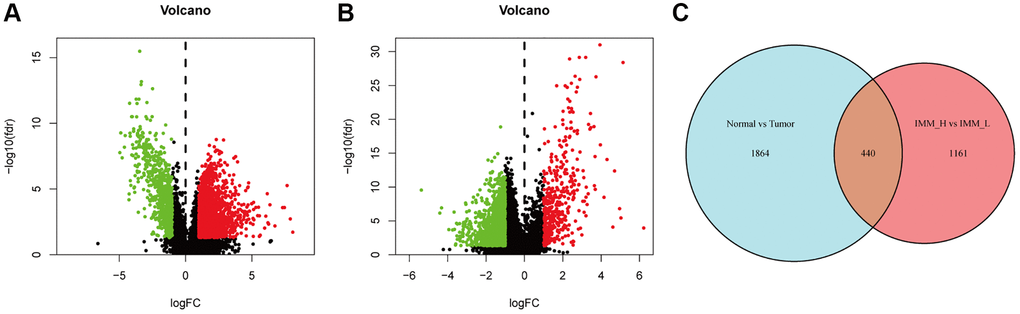
A total of 405 bladder cancer patients from the TCGA-BLCA with 430 transcriptome profiles were obtained (Normal = 19, Tumor = 411). The transcriptome profiles of these samples were conducting ssGSEA analysis to evaluate the immune infiltration and immune related functions. Notably, there were 29 immunological marker gene sets, including immunological cell subtypes, pathways or functions associated with immune, have been used to perform the ssGSEA analysis. Then the samples were clustered into two groups according to the immune infiltration results, named accordingly, high immune cell infiltration cluster (Immunity H, n = 315) and low immune cell infiltration cluster (Immunity L, n = 96) (Figure 1A). The box chart also showed a significant higher Immune Score, Stromal Score, and ESTIMATE score in Immunity H group, while the Immunity L group showed a higher Tumor Purity (Figure 1B). Meanwhile, the expression values of CD274 (PD-L1) and HLA family was discovered significantly higher in Immunity H group than that in Immunity L group (Figure 1C and 1D, p < 0.001). Moreover, the CIBERSORT algorithm was used to verify the reliability of the unsupervised immune subgroups and found that there were higher infiltration of CD8+ T cells and M1 macrophages in the Immunity H group compared to the Immunity L group (Figure 1E). So, all these results showed that this bladder cancer unsupervised grouping performed well in distinguishing the difference of immune infiltration between samples and was suitable for further analysis.
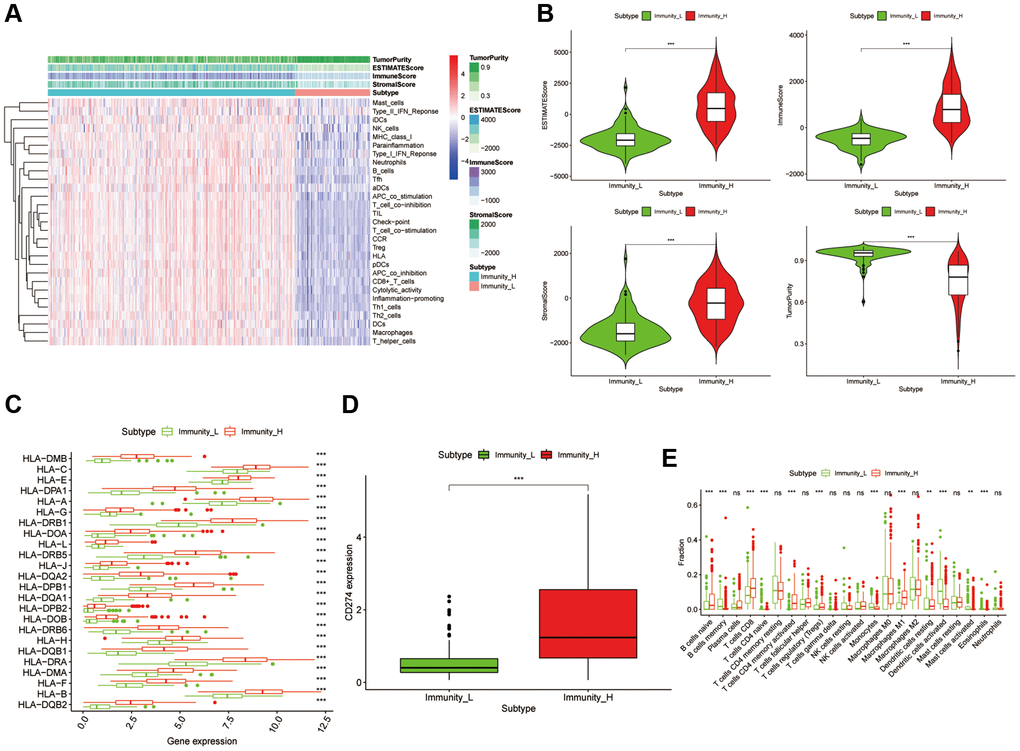
Figure 1. Construction and verification of bladder cancer clustering by immune infiltration. (A) The immune cells were highly infiltrated in the cluster1 group, which was named as the Immunity_H (high immune cell infiltration group), and the low expression one in the cluster2 group was named as the Immunity_L (low immune cell infiltration group). The Tumor Purity, ESTIMATE Score, Immune Score and Stromal Score of each sample gene were also displayed with the grouping information by using ESTIMATE's algorithm. (B) The box-plot showed a statistical difference in Tumor Purity, ESTIMATE Score, Immune Score and Stromal Score between the two groups (p < .01). (C and D) In Immunity_H (red), the expression of HLA family genes and CD274 were all significantly higher than that in the Immunity_L (green) (p < 0.001). (E) The statistical chart showed the proportion difference of each immune cell between the Immunity_H (red) and the Immunity_L (green), after using the CIBERSORT method.
Nomogram and drug response
It seems the risk score is the most weighted factor in the nomogram (Figure 6A), and all of the calibration curves show a consistency between the survival predicted by this nomogram and the actual survival rates (Figure 6B–6D). Besides, we performed drug sensitivity prediction. As shown in the Figure 7A–7E, the ordinate represents IC50, so the smaller the IC50, the more sensitive it is to drugs. So, it is interesting so to see that though patients with high risk scores are associated with a poor prognosis, they showed a more sensitive response to the cisplatin and doxorubicin than patients with low risk scores (Figure 7A and 7B), while they are less sensitive to the methotrexate (Figure 7C). The response to both gemcitabine and vinblastine are no differences (Figure 7D and 7E). Similarly, in our immunotherapy response prediction, the high-risk group showed significantly better response to anti-CTLA4 treatment, while the low-risk patients were more sensitive to PD1 monoclonal antibody (Figure 7F).
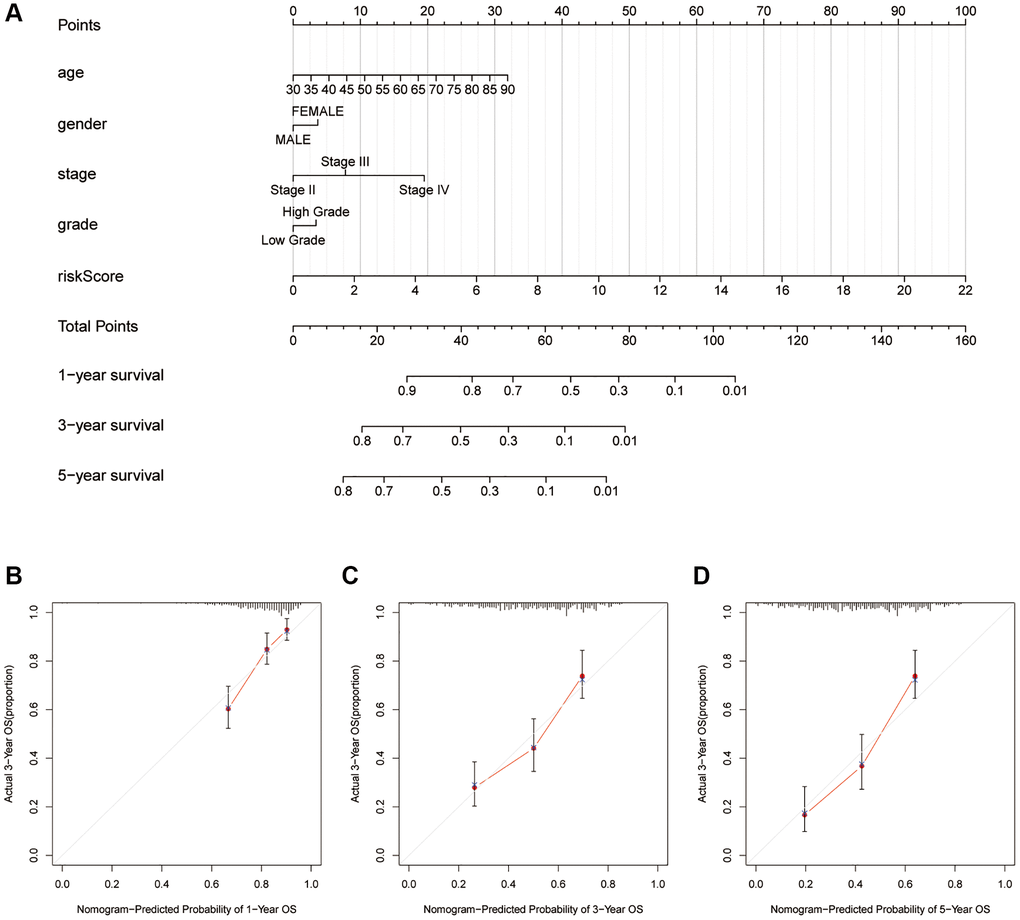
Figure 6. Quantitively analyze the overall survival of patients with bladder cancer. (A) Nomogram considering risk score and several common-used clinicopathological factors. Calibration curves for 1-yaer (B), 3-years (C), 5-years (D).
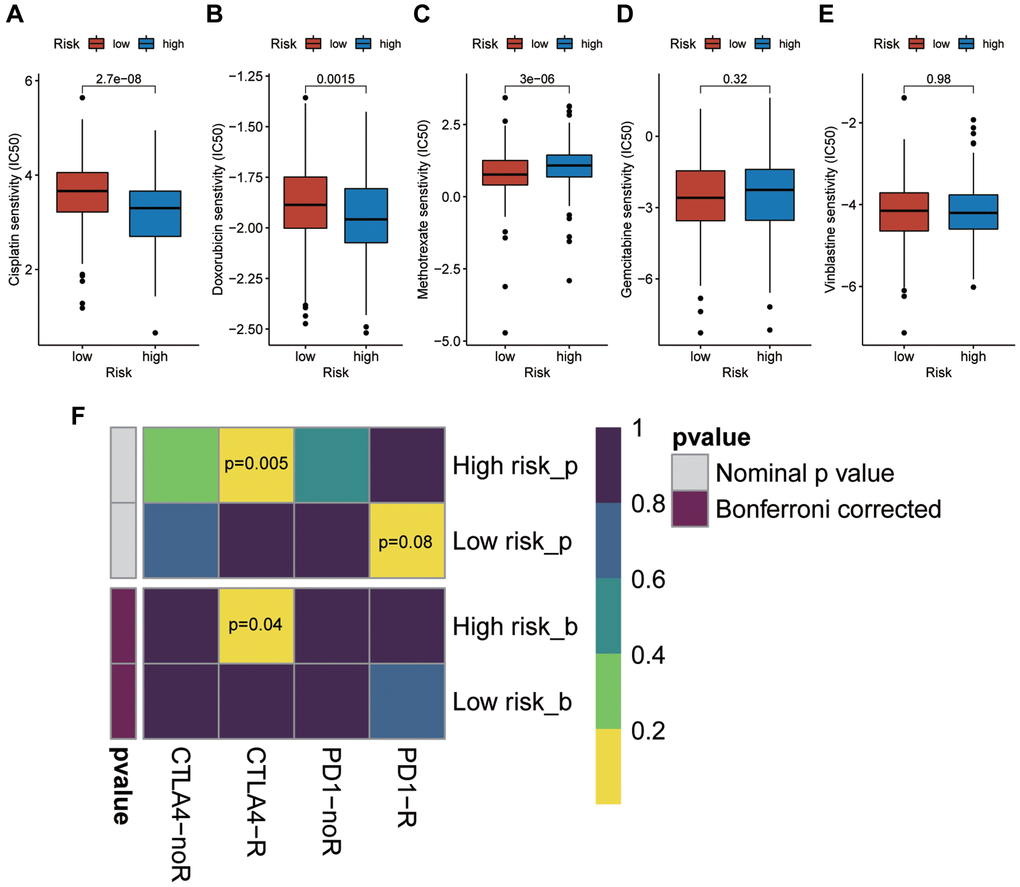
Figure 7. Prediction of the sensitivity of high risk patients to chemotherapy drugs and immunotherapy. (A) Cisplatin. (B) Doxorubicin. (C) Methotrexate. (D) Gemcitabine. (E) Vinblastine. (F) Immune checkpoint receptor: PD1-R, PD1-noR, CTLA4-R, ctal4-noR.
Discussion
BC is one of the common urinary tumor accounts for high prevalence and recurrence rate [23]. BC tissue consists of cancer cells and other stromal cells and immune cells. Among them, immune cells have been proved functioned in tumor progression and prognosis [9]. Besides, TNM staging system cannot distinguish the survival and treatment response of patients accurately that it only considered the anatomical information and ignore the significant role of genetic background, [24]. Thus, scientists devoted to identifying and developing molecular biomarkers for the diagnosis and prognosis in cancer patients [25, 26]. There is significant meaning for discovering the mechanism of the progression in bladder cancer, and the treatment and prognosis of it. Our study focuses more on the immune infiltration related lncRNAs and their interaction with the immune cells. In this study, we identified and verified a 12 immune-infiltration-related lncRNAs as prognostic signature in patients with bladder cancer.
We firstly constructed an unsupervised bladder cancer grouping, dividing into high and low immune infiltration clusters. Then we discovered that there were significant differences in Tumor Purity, ESTIMATE Score, Immune Score, and Stromal Score between these two clusters. Interestingly, the expression of HLA and CD274, also the algorithm of CIBERSORT verified the reliability of this unsupervised immune cluster.
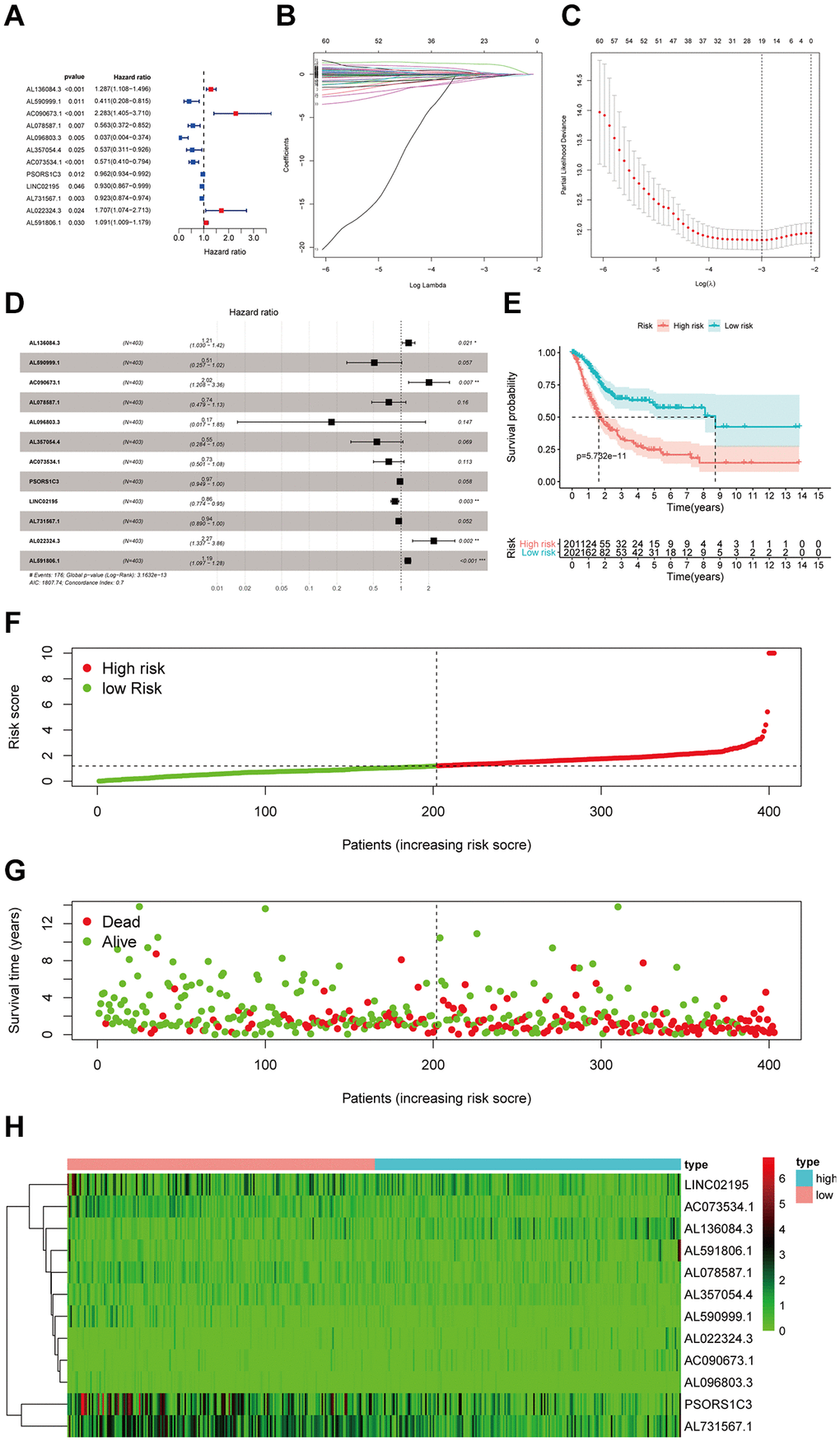
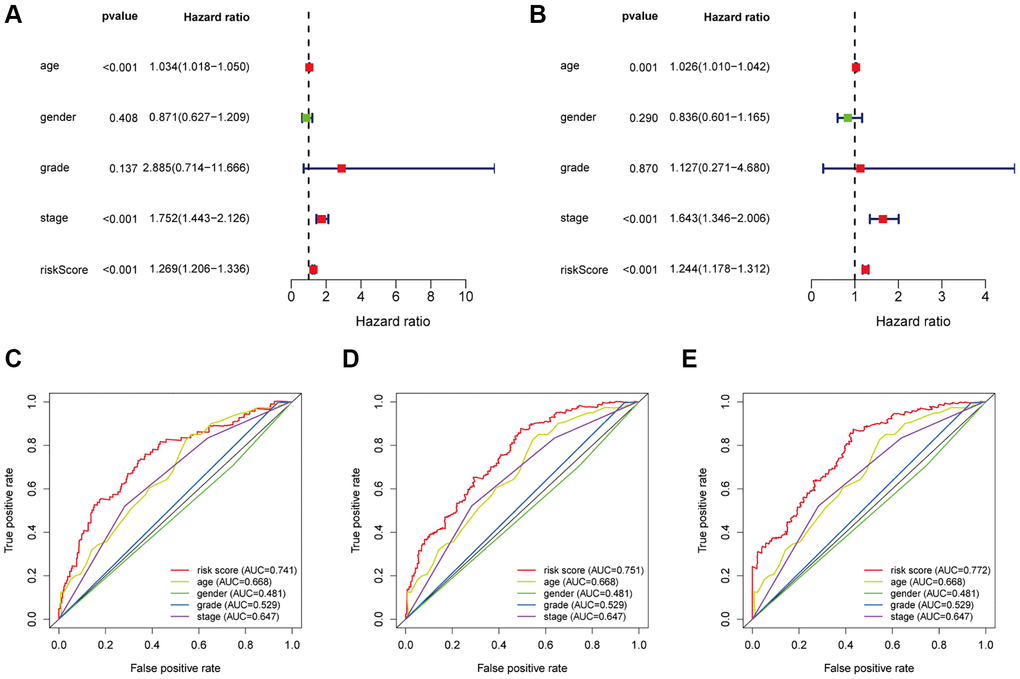
In recent years, under the deep studies in transcriptome sequencing, we have known that around 80% of the transcripts in human genome are noncoding genes, such as lncRNAs, miRNAs, circRNAs, and tsRNAs. Among them, lncRNAs were shown to be associated with the progression, prognosis of bladder cancer [19]. In this study, we identified 12 immune-infiltration-related lncRNAs associated with the prognosis of patients with bladder cancer. Among them, LINC02195 is reported as a favorable prognostic marker in head and neck squamous cell carcinoma [27]. Also, lncRNA PSORS1C3 is discovered expressed regulated by the expression of transcription factor OCT4 in non-pluripotent cells [28]. Having confirmed the significant role of these immune-infiltration related lncRNAs in several biological processes, we constructed the prognostic signature and verified the efficacy of this signature by the univariate and multivariate Cox analysis. Notably, time dependent ROC curves (for 1, 3, 5 years) were plotted to compare the prognosis efficacy between this signature with other common-used clinical factors of patients with bladder cancer, including age, gender, pathological grade and stage.
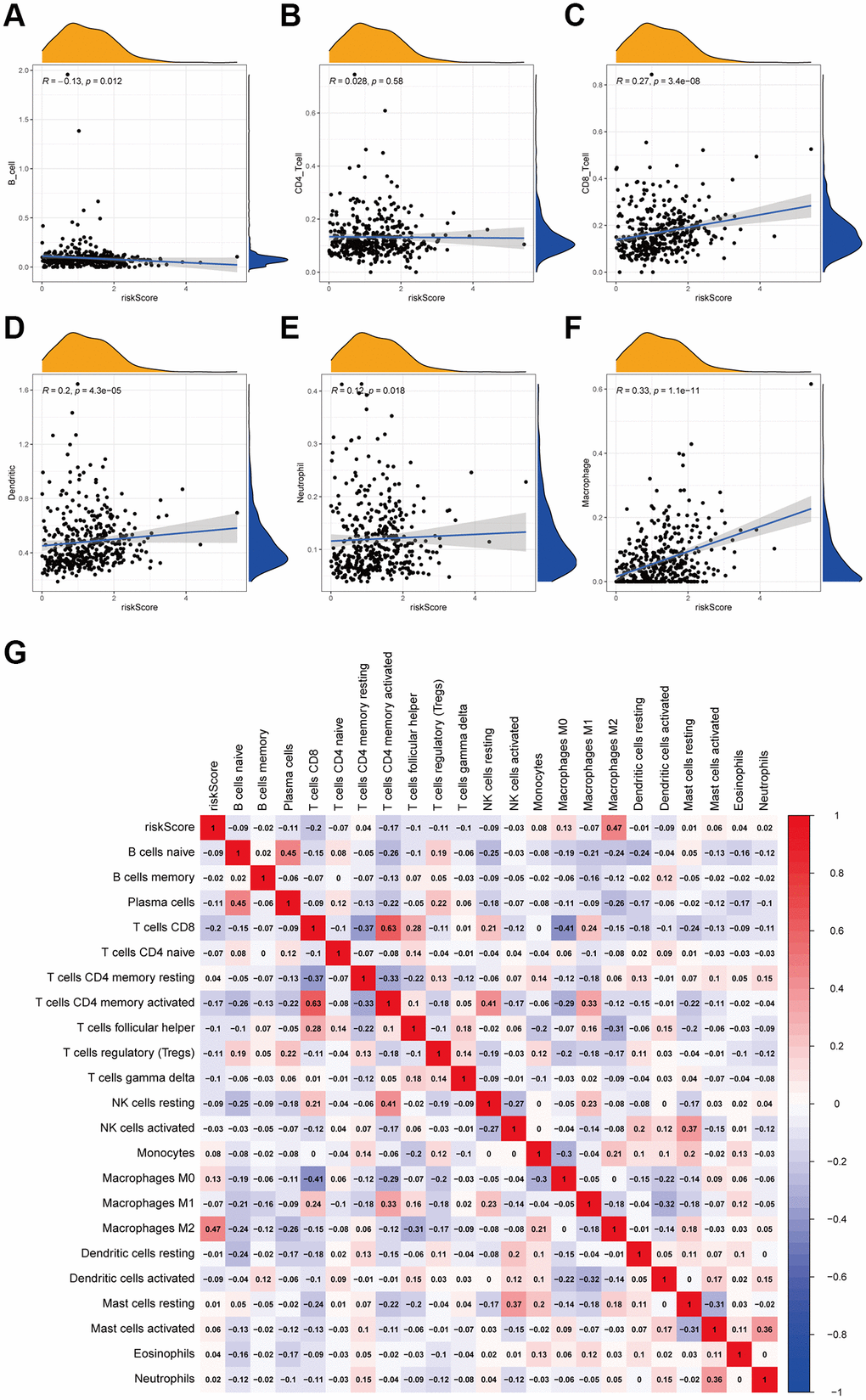
Several studies suggested that tumor-infiltrating lymphocytes are associated with tumor recurrence, progression, and drug response [29, 30]. And it was found that tumor-infiltrating immune cells hold a high infiltration proportion in several types of cancer, for example, breast cancer and skin melanoma [20, 31]. Moreover, the immune infiltration are the main targets of the immunotherapy [32]. In this study, we found that the B cells were significantly negatively-correlated with risk score, and M2 macrophages was positively. B cells, regarded as effector cells of anti-tumor cellular immunity, low infiltration in tumor tissue caused a poor prognosis. On the other hand, M2 macrophages could enhance cell growth [33]. Therefore, these results revealed two potential mechanisms causing worse prognosis in high-risk patients and indicated the potential therapeutic targets in patients with bladder cancer.
In addition, we analyzed the sensitivity of high-risk patients to chemotherapeutic drugs. Interestingly, we found that high risk patients are highly sensitive to cisplatin and doxorubicin. Similarly, these patients were also more sensitive to anti-CTLA4 immunotherapy. We speculate that this may be related to different immune cell infiltration and immune checkpoint expression between high and low risk groups. Therefore, although high risk patients showed a poorer prognosis, these results could provide us with new ideas for targeted treatment among them.
Inevitably, there are some limitations in our research that should be pointed out. Firstly, the 12-lncRNA prognostic signature was only obtained and validated in the TCGA dataset. Secondly, more patient datasets are supposed to verify the performance of the 12-lncRNA prognostic signature. Besides, all the findings need to be verified by more analysis in order to increase authenticity.
In conclusion, our study identified a novel twelve-immune infiltration-related lncRNA signature for bladder cancer. We also found that different score-based groups showed different immune infiltration. These findings may reveal a potential target for the prognostic evaluation of patients with bladder cancer and provide more ideas for further studies on tumor immunity in bladder cancer.
Materials and Methods
Data sources
TCGA_BLCA dataset were retrieved and downloaded in the fragments per kilobase of per million format (FPKM) from the TCGA database (https://portal.gdc.cancer.gov/), the corresponding clinical information containing age, gender, survival status, overall survival, pT stage, pN stage, pM stage, and AJCC stage were also downloaded from TCGA database. A total of 403 bladder cancer patients were enrolled in this research (Table 2). Notably, we obtained annotation gene sets consisting of 29 immune related gene sets considering both immune cells and immune related pathways or functions. Then we performed ssGSEA analysis to emphasize the integrative immune cells, immune related pathways, and immune related functions of each bladder cancer samples by the R package “GSVA”. Following this, all the bladder cancer samples were unsupervised clustered and divided into two clusters defined as high/low immune infiltration group according to the ssGSEA results.
Table 2. Clinical characteristics of the BLCA patients.
| Overall | High risk | Low risk | p | |
| N | 403 | 201 | 202 | |
| Age (mean (SD)) | 68.06 (10.60) | 69.54 (10.05) | 66.58 (10.94) | 0.005 |
| Gender = Female/Male (%) | 105/298 (26.1/73.9) | 60/141 (29.9/70.1) | 45/157 (22.3/77.7) | 0.106 |
| Grade (%) | <0.001 | |||
| High Grade | 380 (94.3) | 197 (98.0) | 183 (90.6) | |
| Low Grade | 20 (5.0) | 1 (0.5) | 19 (9.4) | |
| Unknown | 3 (0.7) | 3 (1.5) | 0 (0.0) | |
| Stage (%) | <0.001 | |||
| Stage I | 1 (0.2) | 0 (0.0) | 1 (0.5) | |
| Stage II | 127 (31.5) | 44 (21.9) | 83 (41.1) | |
| Stage III | 138 (34.2) | 74 (36.8) | 64 (31.7) | |
| Stage IV | 133 (33.0) | 82 (40.8) | 51 (25.2) | |
| Unknown | 4 (1.0) | 1 (0.5) | 3 (1.5) | |
| T (%) | 0.001 | |||
| T0 | 1 (0.2) | 0 (0.0) | 1 (0.5) | |
| T1 | 3 (0.7) | 1 (0.5) | 2 (1.0) | |
| T2 | 37 (9.2) | 8 (4.0) | 29 (14.4) | |
| T2a | 25 (6.2) | 9 (4.5) | 16 (7.9) | |
| T2b | 56 (13.9) | 27 (13.4) | 29 (14.4) | |
| T3 | 42 (10.4) | 23 (11.4) | 19 (9.4) | |
| T3a | 69 (17.1) | 32 (15.9) | 37 (18.3) | |
| T3b | 80 (19.9) | 49 (24.4) | 31 (15.3) | |
| T4 | 10 (2.5) | 9 (4.5) | 1 (0.5) | |
| T4a | 43 (10.7) | 27 (13.4) | 16 (7.9) | |
| T4b | 5 (1.2) | 4 (2.0) | 1 (0.5) | |
| TX | 1 (0.2) | 0 (0.0) | 1 (0.5) | |
| Unknown | 31 (7.7) | 12 (6.0) | 19 (9.4) | |
| M (%) | 0.015 | |||
| M0 | 193 (47.9) | 82 (40.8) | 111 (55.0) | |
| M1 | 11 (2.7) | 8 (4.0) | 3 (1.5) | |
| MX | 197 (48.9) | 110 (54.7) | 87 (43.1) | |
| Unknown | 2 (0.5) | 1 (0.5) | 1 (0.5) | |
| N (%) | 0.028 | |||
| N0 | 234 (58.1) | 104 (51.7) | 130 (64.4) | |
| N1 | 46 (11.4) | 31 (15.4) | 15 (7.4) | |
| N2 | 75 (18.6) | 43 (21.4) | 32 (15.8) | |
| N3 | 7 (1.7) | 5 (2.5) | 2 (1.0) | |
| NX | 36 (8.9) | 15 (7.5) | 21 (10.4) | |
| Unknown | 5 (1.2) | 3 (1.5) | 2 (1.0) | |
| RiskScore (median [IQR]) | 1.18 [0.72, 1.76] | 1.76 [1.50, 2.14] | 0.72 [0.43, 0.95] | <0.001 |
Characteristics of the immune grouping
Having obtained the immune grouping of each sample, we would like to verify the effective ness of this immune cluster. Thus, we firstly conducted ESTIMATE analysis to emphasize the corresponding immune score, stromal score, estimate score, and tumor purity of each sample according to their transcriptional expression. Then, all these immune related scores were compared between these two immune groups. Besides, the gene expression of immune checkpoint including human leukocyte antigen (HLA) and CD274 (PD-L1) were compared between high immune cell infiltration group (Immunity H) and low immune cell infiltration group (Immunity L) to further verifying the effectiveness of the immune infiltration groups. Moreover, CIBERSORT algorithm was performed to estimate the detailed immune cell infiltration status of each sample, and Wilcoxon test was carried out to investigate the differential immune cell infiltration between these two immune infiltration groups.
Correlation between prognostic signature and detailed immune infiltration
Here we took two differential methods to estimate the immune cell infiltration of each sample, and they were TIMER and CIBERORT. Among them, the immune cells calculated by TIMER was downloaded from the TIMER database (https://cistrome.shinyapps.io/timer/). The CIBERSORT results were calculated by the CIBERSORT algorithm. Following these two methods, PEARSON correlation test was carried out to further investigating the correlation between risk score and these detailed immune cells infiltration.
Nomogram and drug response
To emphasize the overall survival of patients with bladder cancer more quantitatively, we re-checked the corresponding clinical data of each patient and assembled a nomogram considering risk score and several common-used clinicopathological factors including age, gender, stage, and grade. Calibration curves for 1-, 3-, 5-years were also plotted to examine the accuracy of the nomogram. Finally, we separately predicted the drug response to chemotherapy by R package “ProPhetic” and the drug response to immunotherapy by submap algorithm [34]. Then compared the drug response to both chemotherapy and immunotherapy between high and low risk patients.
Statistical analysis
All statistical analysis was applied by R program version 4.0.2. Mean ± standard deviation was applied to describe the distribution of the continuous variables following normal distribution while the median (range) was used for continuous variables following abnormal distribution. Counts and percentages were used to describe the distribution of categorical variables. Two-tailed p < 0.05 was considered with statistical significance.
Data availability
Source data of this study were derived from the public repositories, as indicated in the section of “Materials and Methods” of the manuscript. And all data that support the findings of this study are available from the corresponding author upon reasonable request.
Supplementary Materials
Author Contributions
XQD and LCQ: design, analysis and interpretation of data, drafting of the manuscript, critical revision of the manuscript; XQD: statistical analysis; XQD, LCQ, LJL, SJX, XJZ, LZ, HJ and WSG: methodology; LCQ, XQD, LZ, HJ and WSG: project administration; LCQ and XQD: Writing (original draft); LCQ, XQD, LJL, SJX, XJZ, LZ, HJ and WSG: Writing (review and editing).
Acknowledgments
We thank all the R programming package developer.
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
Funding
This research is supported by grants from the National Natural Science Foundation of China (NSFC, 81772729) and grants from Natural Science Foundation of Hubei Province (ZRMS2020002466).
References
- 1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492 [PubMed]
- 2. Feng RM, Zong YN, Cao SM, Xu RH. Current cancer situation in China: good or bad news from the 2018 Global Cancer Statistics? Cancer Commun (Lond). 2019; 39:22. https://doi.org/10.1186/s40880-019-0368-6 [PubMed]
- 3. Qiu H, Hu X, He C, Yu B, Li Y, Li J. Identification and Validation of an Individualized Prognostic Signature of Bladder Cancer Based on Seven Immune Related Genes. Front Genet. 2020; 11:12. https://doi.org/10.3389/fgene.2020.00012 [PubMed]
- 4. Cavallo F, De Giovanni C, Nanni P, Forni G, Lollini PL. 2011: the immune hallmarks of cancer. Cancer Immunol Immunother. 2011; 60:319–26. https://doi.org/10.1007/s00262-010-0968-0 [PubMed]
- 5. Zitvogel L, Tesniere A, Kroemer G. Cancer despite immunosurveillance: immunoselection and immunosubversion. Nat Rev Immunol. 2006; 6:715–27. https://doi.org/10.1038/nri1936 [PubMed]
- 6. Balkwill F, Mantovani A. Inflammation and cancer: back to Virchow? Lancet. 2001; 357:539–45. https://doi.org/10.1016/S0140-6736(00)04046-0 [PubMed]
- 7. Harao M, Forget MA, Roszik J, Gao H, Babiera GV, Krishnamurthy S, Chacon JA, Li S, Mittendorf EA, DeSnyder SM, Rockwood KF, Bernatchez C, Ueno NT, et al. 4-1BB-Enhanced Expansion of CD8+ TIL from Triple-Negative Breast Cancer Unveils Mutation-Specific CD8+ T Cells. Cancer Immunol Res. 2017; 5:439–45. https://doi.org/10.1158/2326-6066.CIR-16-0364 [PubMed]
- 8. Liu K, Zhao K, Wang L, Sun E. The prognostic values of tumor-infiltrating neutrophils, lymphocytes and neutrophil/lymphocyte rates in bladder urothelial cancer. Pathol Res Pract. 2018; 214:1074–80. https://doi.org/10.1016/j.prp.2018.05.010 [PubMed]
- 9. Sharma P, Shen Y, Wen S, Yamada S, Jungbluth AA, Gnjatic S, Bajorin DF, Reuter VE, Herr H, Old LJ, Sato E. CD8 tumor-infiltrating lymphocytes are predictive of survival in muscle-invasive urothelial carcinoma. Proc Natl Acad Sci U S A. 2007; 104:3967–72. https://doi.org/10.1073/pnas.0611618104 [PubMed]
- 10. El-Dairi M, House RJ. Optic nerve hypoplasia. Handbook of Pediatric Retinal OCT and the Eye-Brain Connection. Elsevier. 2020; 285–7. https://doi.org//10.1016/B978-0-323-60984-5.00062-7
- 11. Sorigue M, Ribera JM, Sancho JM. The clinical dilemma of grade 3 follicular lymphoma. Ann Oncol. 2016; 27:1974. https://doi.org/10.1093/annonc/mdw284 [PubMed]
- 12. Kang BW, Seo AN, Yoon S, Bae HI, Jeon SW, Kwon OK, Chung HY, Yu W, Kang H, Kim JG. Prognostic value of tumor-infiltrating lymphocytes in Epstein-Barr virus-associated gastric cancer. Ann Oncol. 2016; 27:494–501. https://doi.org/10.1093/annonc/mdv610 [PubMed]
- 13. Adams S, Gray RJ, Demaria S, Goldstein L, Perez EA, Shulman LN, Martino S, Wang M, Jones VE, Saphner TJ, Wolff AC, Wood WC, Davidson NE, et al. Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin Oncol. 2014; 32:2959–66. https://doi.org/10.1200/JCO.2013.55.0491 [PubMed]
- 14. Xue Y, Tong L, LiuAnwei Liu F, Liu A, Zeng S, Xiong Q, Yang Z, He X, Sun Y, Xu C. Tumor-infiltrating M2 macrophages driven by specific genomic alterations are associated with prognosis in bladder cancer. Oncol Rep. 2019; 42:581–94. https://doi.org/10.3892/or.2019.7196 [PubMed]
- 15. Stefanski CD, Keffler K, McClintock S, Milac L, Prosperi JR. APC loss affects DNA damage repair causing doxorubicin resistance in breast cancer cells. Neoplasia. 2019; 21:1143–50. https://doi.org/10.1016/j.neo.2019.09.002 [PubMed]
- 16. Michea P, Noël F, Zakine E, Czerwinska U, Sirven P, Abouzid O, Goudot C, Scholer-Dahirel A, Vincent-Salomon A, Reyal F, Amigorena S, Guillot-Delost M, Segura E, Soumelis V. Adjustment of dendritic cells to the breast-cancer microenvironment is subset specific. Nat Immunol. 2018; 19:885–97. https://doi.org/10.1038/s41590-018-0145-8 [PubMed]
- 17. Cai P, Otten ABC, Cheng B, Ishii MA, Zhang W, Huang B, Qu K, Sun BK. A genome-wide long noncoding RNA CRISPRi screen identifies PRANCR as a novel regulator of epidermal homeostasis. Genome Res. 2020; 30:22–34. https://doi.org/10.1101/gr.251561.119 [PubMed]
- 18. Shen Y, Peng X, Shen C. Identification and validation of immune-related lncRNA prognostic signature for breast cancer. Genomics. 2020; 112:2640–6. https://doi.org/10.1016/j.ygeno.2020.02.015 [PubMed]
- 19. Quan J, Pan X, Zhao L, Li Z, Dai K, Yan F, Liu S, Ma H, Lai Y. LncRNA as a diagnostic and prognostic biomarker in bladder cancer: a systematic review and meta-analysis. Onco Targets Ther. 2018; 11:6415–24. https://doi.org/10.2147/OTT.S167853 [PubMed]
- 20. Yu WD, Wang H, He QF, Xu Y, Wang XC. Long noncoding RNAs in cancer-immunity cycle. J Cell Physiol. 2018; 233:6518–23. https://doi.org/10.1002/jcp.26568 [PubMed]
- 21. Wang Y, Yang X, Sun X, Rong L, Kang M, Wu P, Ji X, Lin R, Huang J, Xue Y, Fang Y. Bone marrow infiltrated Lnc-INSR induced suppressive immune microenvironment in pediatric acute lymphoblastic leukemia. Cell Death Dis. 2018; 9:1043. https://doi.org/10.1038/s41419-018-1078-8 [PubMed]
- 22. Zhou M, Zhang Z, Bao S, Hou P, Yan C, Su J, Sun J. Computational recognition of lncRNA signature of tumor-infiltrating B lymphocytes with potential implications in prognosis and immunotherapy of bladder cancer. Brief Bioinform. 2021; 22:bbaa047. https://doi.org/10.1093/bib/bbaa047 [PubMed]
- 23. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020; 70:7–30. https://doi.org/10.3322/caac.21590 [PubMed]
- 24. Qu L, Wang ZL, Chen Q, Li YM, He HW, Hsieh JJ, Xue S, Wu ZJ, Liu B, Tang H, Xu XF, Xu F, Wang J, et al. Prognostic Value of a Long Non-coding RNA Signature in Localized Clear Cell Renal Cell Carcinoma. Eur Urol. 2018; 74:756–63. https://doi.org/10.1016/j.eururo.2018.07.032 [PubMed]
- 25. Choi W, Ochoa A, McConkey DJ, Aine M, Höglund M, Kim WY, Real FX, Kiltie AE, Milsom I, Dyrskjøt L, Lerner SP. Genetic Alterations in the Molecular Subtypes of Bladder Cancer: Illustration in the Cancer Genome Atlas Dataset. Eur Urol. 2017; 72:354–65. https://doi.org/10.1016/j.eururo.2017.03.010 [PubMed]
- 26. Li X, Wu WK, Xing R, Wong SH, Liu Y, Fang X, Zhang Y, Wang M, Wang J, Li L, Zhou Y, Tang S, Peng S, et al. Distinct Subtypes of Gastric Cancer Defined by Molecular Characterization Include Novel Mutational Signatures with Prognostic Capability. Cancer Res. 2016; 76:1724–32. https://doi.org/10.1158/0008-5472.CAN-15-2443 [PubMed]
- 27. Li H, Xiong HG, Xiao Y, Yang QC, Yang SC, Tang HC, Zhang WF, Sun ZJ. Long Non-coding RNA LINC02195 as a Regulator of MHC I Molecules and Favorable Prognostic Marker for Head and Neck Squamous Cell Carcinoma. Front Oncol. 2020; 10:615. https://doi.org/10.3389/fonc.2020.00615 [PubMed]
- 28. Mirzadeh Azad F, Malakootian M, Mowla SJ. lncRNA PSORS1C3 is regulated by glucocorticoids and fine-tunes OCT4 expression in non-pluripotent cells. Sci Rep. 2019; 9:8370. https://doi.org/10.1038/s41598-019-44827-7 [PubMed]
- 29. Stanton SE, Disis ML. Clinical significance of tumor-infiltrating lymphocytes in breast cancer. J Immunother Cancer. 2016; 4:59. https://doi.org/10.1186/s40425-016-0165-6 [PubMed]
- 30. Fu Q, Chen N, Ge C, Li R, Li Z, Zeng B, Li C, Wang Y, Xue Y, Song X, Li H, Li G. Prognostic value of tumor-infiltrating lymphocytes in melanoma: a systematic review and meta-analysis. Oncoimmunology. 2019; 8:1593806. https://doi.org/10.1080/2162402X.2019.1593806 [PubMed]
- 31. Huang R, Zeng Z, Li G, Song D, Yan P, Yin H, Hu P, Zhu X, Chang R, Zhang X, Zhang J, Meng T, Huang Z. The Construction and Comprehensive Analysis of ceRNA Networks and Tumor-Infiltrating Immune Cells in Bone Metastatic Melanoma. Front Genet. 2019; 10:828. https://doi.org/10.3389/fgene.2019.00828 [PubMed]
- 32. Jiang R, Tang J, Chen Y, Deng L, Ji J, Xie Y, Wang K, Jia W, Chu WM, Sun B. The long noncoding RNA lnc-EGFR stimulates T-regulatory cells differentiation thus promoting hepatocellular carcinoma immune evasion. Nat Commun. 2017; 8:15129. https://doi.org/10.1038/ncomms15129 [PubMed]
- 33. Sharifi L, Nowroozi MR, Amini E, Arami MK, Ayati M, Mohsenzadegan M. A review on the role of M2 macrophages in bladder cancer; pathophysiology and targeting. Int Immunopharmacol. 2019; 76:105880. https://doi.org/10.1016/j.intimp.2019.105880 [PubMed]
- 34. Hoshida Y, Brunet JP, Tamayo P, Golub TR, Mesirov JP. Subclass mapping: identifying common subtypes in independent disease data sets. PLoS One. 2007; 2:e1195. https://doi.org/10.1371/journal.pone.0001195 [PubMed]