Introduction
Bladder urothelial carcinoma (BLCA) is the most common malignant tumor of the urinary system, accounting for 6.6% and 2.1% of the total cancer patients among men and women in the world, respectively [1, 2]. Patients with transitional cell carcinoma account for approximately 90% of all BCLA cases [3]. Despite great strides in radiotherapy, surgery, and adjuvant chemotherapy, the survival outcomes remain poor for BCLA patients, with approximately 30% of the patients diagnosed with advanced muscle-invasive disease [2]. Moreover, the current clinical staging system requires improvement in accurately predicting the prognosis of BCLA patients [4].
Autophagy is an evolutionarily conserved catabolic process, which occurs at basal levels under normal conditions to eliminate worn out cellular organelles and damaged or mis-folded proteins [5]. However, dysregulation of autophagy is implicated in several human diseases, such as cancer [6], neurodegenerative disorders [7], cardiovascular diseases [8], and inflammatory disorders related to infectious diseases [9]. Autophagy is associated with tumor suppression or oncogenesis depending upon the stage of tumor development [10, 11]. Recent studies show that modulation of autophagy improves the sensitivity of BCLA tumors to chemotherapeutic agents [12, 13]. Hence, it is critical to discover autophagy-related biomarkers that can serve as valuable the early diagnostic and prognostic biomarkers for BCLA patients.
Genome sequencing studies show that nearly 90% of the human transcriptome represents non-coding RNA or ncRNA [14]. The long non-coding RNAs (lncRNAs) are a type of ncRNAs with transcripts of >200 nucleotides in length without any protein-coding capacity [15]. LncRNAs regulate important biological functions related to cell growth and survival, genomic imprinting, chromatin modifications, and allosteric regulation of enzyme activities [16]. Furthermore, pathogenesis of several human diseases including several different types of cancers involves dysregulation of specific lncRNAs [17]. Some studies have shown that lncRNAs regulate autophagic functions. For example, Ying et al. demonstrated that downregulation of lncRNA MEG3 promotes proliferation of bladder cancer cells by activating autophagy [18]. Another study shows that lncRNA MALAT1 regulates multi-drug resistance of hepatocellular carcinoma cells by altering autophagy [19]. LncRNA PVT1 promotes in vitro and in vivo pancreatic ductal adenocarcinoma progression by activating autophagy through its regulation of the miR-20a-5p/ULK1 axis [20].
New advances in genome sequencing technology and bioinformatics have helped to identify potential prognostic biomarkers that can predict survival outcomes in cancer patients [21, 22]. Therefore, we postulated that autophagy-related lncRNAs may be valuable prognostic biomarkers for BLCA patients. In this study, we systematically analyzed the relationship between the expression of autophagy-related lncRNAs and the clinicopathological characteristics of 409 BLCA patients from The Cancer Genome Atlas (TCGA) database. We also constructed a prognostic signature based on 5 autophagy-related lncRNAs and evaluated its ability to independently and accurately predict the prognosis of BLCA patients.
Results
Construction of the lncRNA–mRNA co-expression network and functional enrichment analysis
Next, we investigated the potential functions of the 5 autophagy-related lncRNAs in BLCA by constructing the lncRNA-mRNA co-expression network using Cytoscape. The lncRNA-mRNA co-expression network contained 77 lncRNA-mRNA pairs based on the threshold parameters, Pearson correlation coefficient |R| > 0.3 and P < 0.05 (Figure 6A). Among these, 49 mRNAs significantly correlated with the 5 lncRNAs in the prognostic signature. The Sankey diagram showed the relationship between the 49 mRNAs and 5 lncRNAs (risk/protective) (Figure 6B). The top three GO terms for the biological processes were autophagy, process utilizing autophagic mechanism, and macroautophagy (Figure 6C). The top three GO terms for the cellular components were cytosolic part, PML body, and the nuclear envelope (Figure 6D). The top three GO terms for molecular functions were protein serine/threonine kinase activity, ubiquitin protein ligase binding, and ubiquitin−like protein ligase binding (Figure 6E). KEGG pathway analysis confirmed that autophagy was the most significantly enriched pathway (Figure 6F).
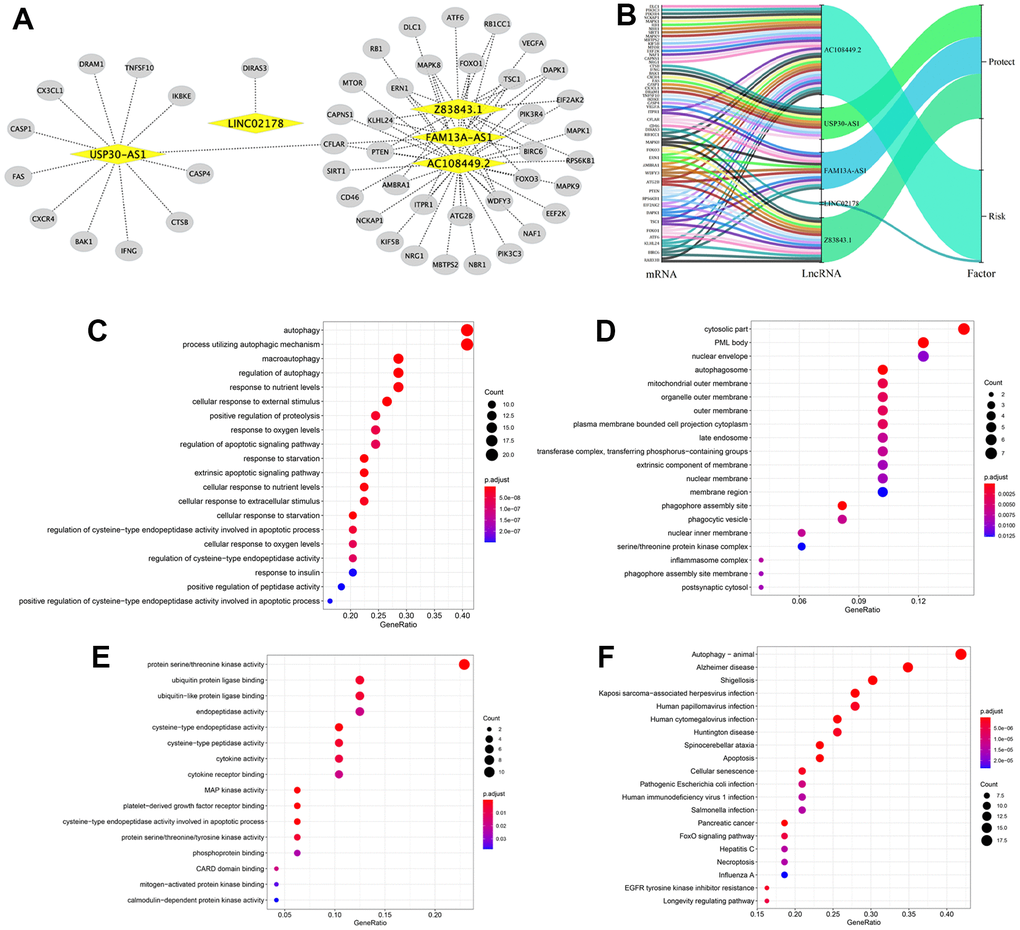
Figure 6. Construction of the autophagy-related lncRNA–mRNA co-expression network and functional enrichment analyses. (A) Diagrammatic representation of the autophagy-related lncRNA–mRNA network shows 77 lncRNA-mRNA co-expression pairs formed between 5 autophagy-related lncRNAs and 49 mRNAs. The yellow circles correspond to autophagy-related lncRNAs, and the gray diamonds correspond to the mRNAs. Every edge represents a co-expression relationship between an lncRNA and an mRNA in the context of BCLA. (B) The Sankey diagram shows the connection degree between the 49 mRNAs and 5 autophagy-related lncRNAs (risk/protective). (C–E) Gene Ontology (GO) analysis results show the enriched (C) biological processes, (D) cell components and (E) molecular functions associated with the mRNAs that co-express with the 5 autophagy-related lncRNAs. (F) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis results shows the enriched signaling pathways associated with the mRNAs that co-express with the 5 autophagy-related lncRNAs.
Gene set enrichment analysis
GSEA results showed that the altered genes in the high-risk BCLA patients belonged to pathways related to autophagy and cancer, WNT signaling pathway, renal cell carcinoma, TGF-βsignaling pathway, VEGF signaling pathway, ERBB signaling pathway, PPAR signaling pathway, MAPK signaling pathway, P53 signaling pathway, mTOR signaling pathway, endocytosis, RNA degradation and ubiquitin-mediated proteolysis (Figure 7A). Immunoregulatory pathways against cancer were significantly enriched in the low-risk group, including pathways related to antigen processing and presentation, natural killer (NK) cell-mediated cytotoxicity, T cell receptor (TCR) signaling, chemokine signaling and B cell receptor (BCR) signaling (Figure 7B). This suggested that activation of pathways regulation immune function in the low-risk group may contribute to positive prognosis or longer survival outcomes. The top 10 KEGG pathways in the high-risk and low-risk groups based on GSEA are shown in Figure 7C and 7D. These results suggested that a high prognostic signature risk score correlates with autophagy and cancer, whereas low prognostic signature risk score correlates with enhanced immune function. These data provided valuable insights for future investigations into potential individualized treatments for BLCA patients belonging to different risk groups.
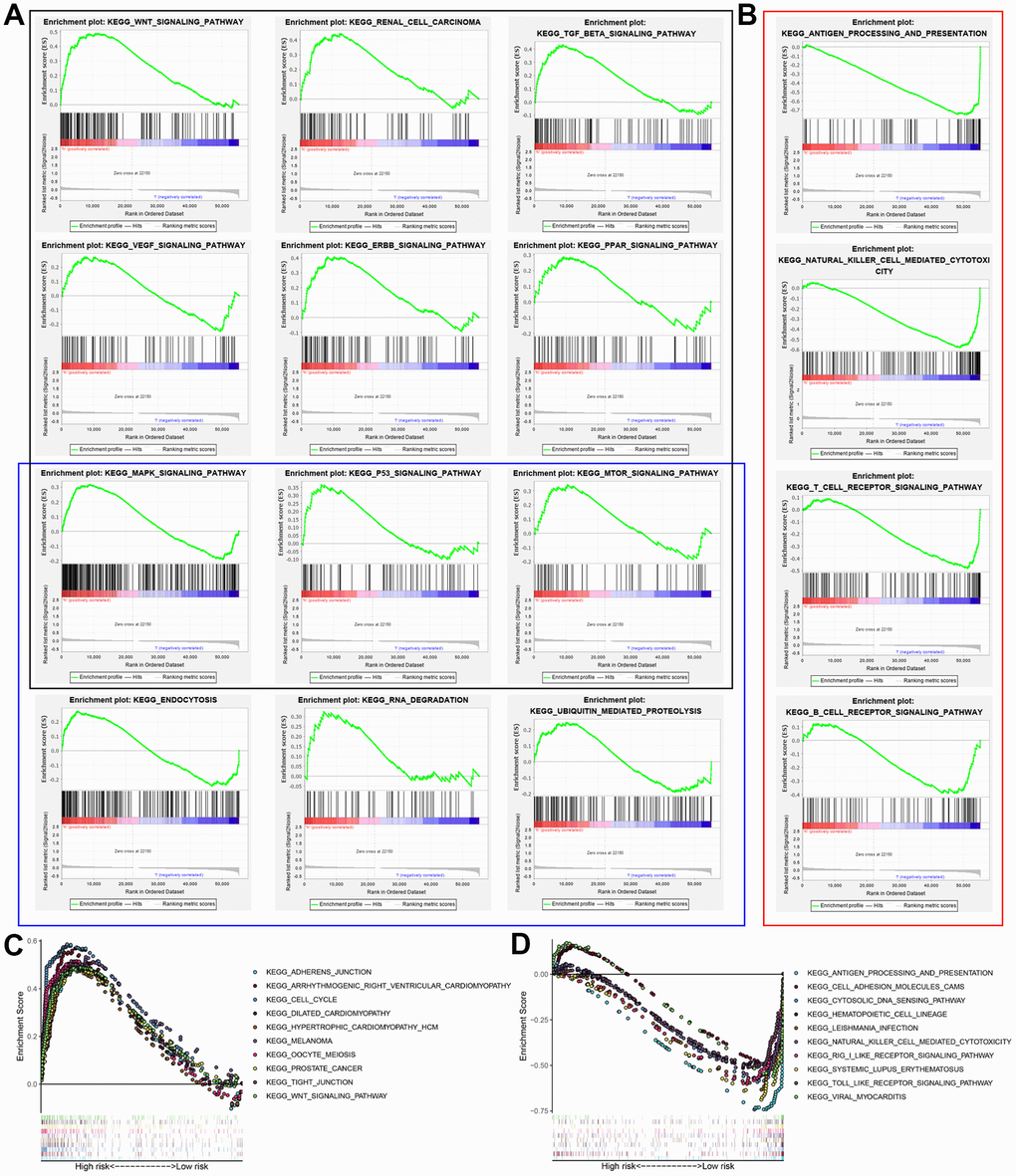
Figure 7. Gene set enrichment analysis (GSEA) of high-risk and low-risk BCLA patients based on the autophagy-related lncRNA prognostic signature. (A) GSEA results show significant enrichment of cancer- and autophagy-related signaling pathways in the high-risk BCLA patients. The black and blue boxes correspond to cancer-related and autophagy-related KEGG signaling pathways, respectively. (B) GSEA results show significant enrichment of immunoregulatory signaling pathways in the low-risk BCLA patients. (C, D) The top 10 KEGG signaling pathways in the (C) high-risk and (D) low-risk BCLA patients.
Discussion
The most common malignancy of the urinary system is BLCA, whose incidence rates are constantly increasing worldwide [1, 3]. The prognosis of BCLA patients is poor because of late diagnosis and high rate of therapeutic resistance [24]. The role of autophagy in tumorigenesis has been reported for several cancers, including BLCA [25]. Therefore, autophagy-related biomarkers are potential diagnostic biomarkers and therapeutic targets for BCLA patients. Previous studies have focused on the role of specific autophagy-related genes in BCLA progression [26].
LncRNAs are a new class of non-coding RNA molecules that regulate cancer cell growth, progression, and survival [27]. Hence, they are potential biomarkers that can predict cancer risk and survival outcomes. In this study, we systematically analyzed the prognostic prediction accuracy of autophagy-related lncRNAs in BCLA using bioinformatics and statistical tools.
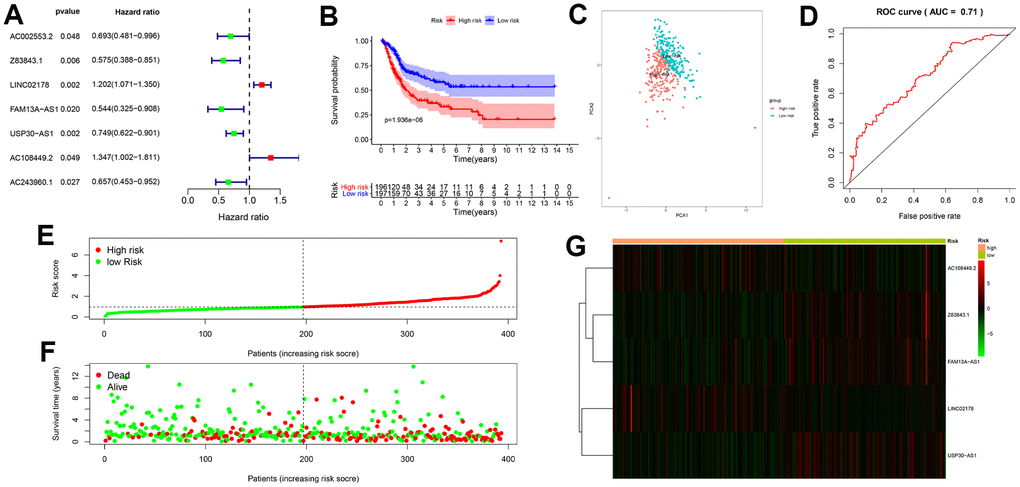
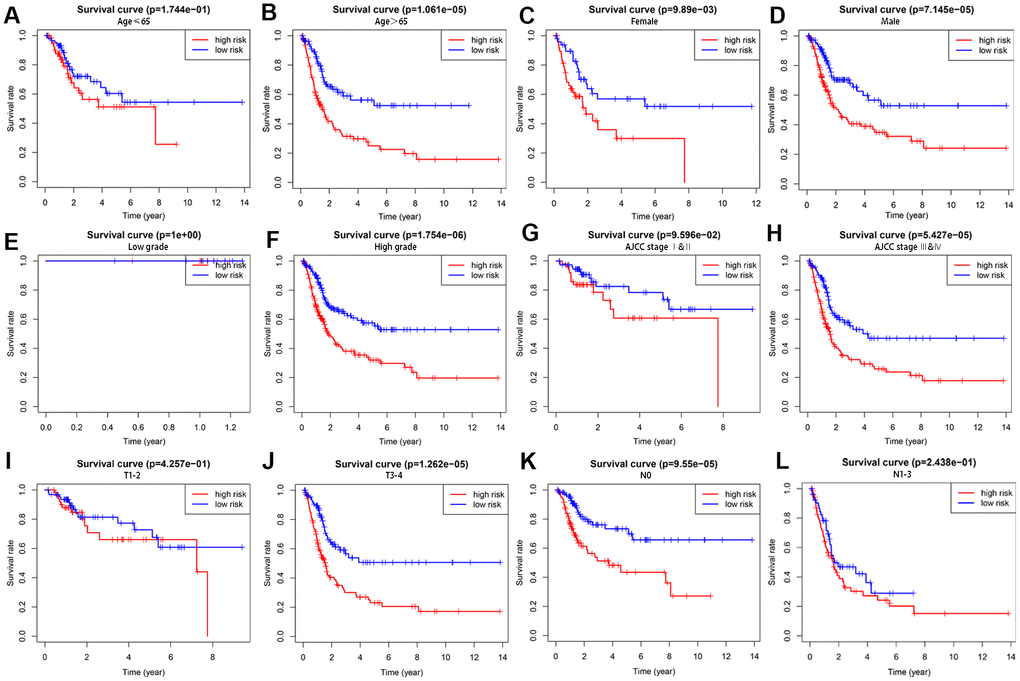
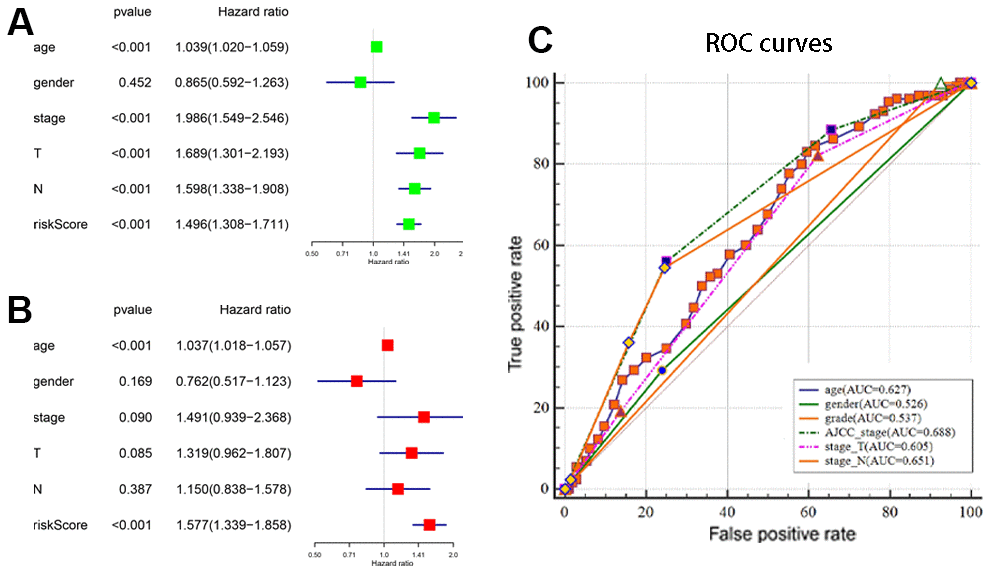
We first identified 7 autophagy-related lncRNAs that significantly correlated with OS based on the univariate Cox regression analysis of the expression of autophagy-related lncRNAs in the BCLA patient samples from the TCGA database. Furthermore, 5 autophagy-related lncRNAs, Z83843.1, LINC02178, FAM13A−AS1, USP30−AS1 and AC108449.2 were selected to construct a prognostic signature based on their performance in the multivariate Cox regression analysis. The risk score of each BCLA patient was calculated according to the expression of the five autophagy-related lncRNAs in the prognostic signature and patients were classified into high- and low- risk groups based on the median risk score. BCLA patients with high-risk scores showed shorter survival times compared to those with low-risk scores. Principal components analysis (PCA) based on the confirmed five autophagy-related lncRNAs showed two significantly different distribution patterns between high-risk and low-risk groups. ROC curve analysis validated the prognostic accuracy of the autophagy-related lncRNA prognostic signature in the BLCA patients. The risk score based on the autophagy-related lncRNA prognostic signature was an independent prognostic factor based on multivariate Cox regression analysis. Stratified correlation analysis showed that the autophagy-related lncRNA prognostic signature accurately predicted survival outcomes for the high- and low-risk BCLA patients.
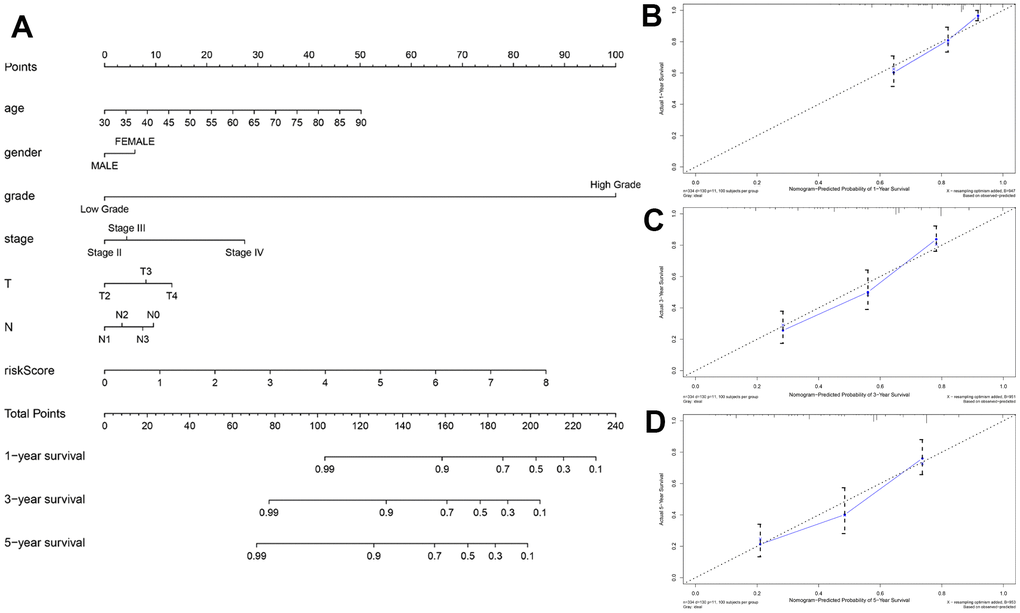
The autophagy-related lncRNA prognostic signature performed more reliably than the other traditional clinical indicators in prognostic prediction. A nomogram is an effective and reliable clinical tool to predict survival of cancer patients [28]. Therefore, we developed a robust nomogram consisting of several clinical variables (age, gender, grade, AJCC stage, T stage and N stage) and the risk scores based on the autophagy-related lncRNA prognostic signature to improve prognostic prediction of BCLA patients. Older patients (age ≥ 60 years) and those with higher tumor grades and advanced stages are usually associated with worse cancer prognosis, which is consistent with our results. Moreover, calibration plots demonstrated that the actual and predicted 1-, 3-, and 5-year survival rates based on the nomogram were similar. Overall, the autophagy-related lncRNA prognostic signature accurately predicts survival outcomes of BLCA patients in our study and shows great potential for clinical applications, including individualized prognosis and therapy.
Autophagy is a highly conserved intracellular catabolic process involved in the phagocytosis and degradation of abnormal organelles, proteins and pathogens through the lysosomal pathway [29]. The role of autophagy in cancer is controversial because it can play both tumor suppressor and oncogenic functions [30]. During early stages of tumor development, autophagy-related cell death can suppress tumor progression; autophagic dysregulation can also induce genomic instability and necrosis-induced inflammation, both of which promote tumor growth [31]. Conversely, autophagy sustains tumor metabolism, growth, and survival in nutrient-deprived conditions in the tumor microenvironment and contributes to drug resistance during tumor metastasis [32]. Autophagy is regulated by several signaling pathways, including the PI3K/AKT/mTOR signaling pathway [33] and ubiquitin-proteasome system (UPS). [34] In recent years, several lncRNAs have been implicated in the regulation of cell growth and survival by directly targeting autophagy-related genes. For example, Wang et al. reported that lncRNA ATB induced autophagy by enhancing the expression of autophagy-related protein 5 (ATG5) through activation of the Yes-associated protein (YAP) in hepatocellular carcinoma (HCC) cells [35]. We identified the genes whose expression is regulated by each of the 5 autophagy-associated lncRNAs in BLCA and constructed the lncRNA-mRNA co-expression network. GO, KEGG, and GSEA functional enrichment analyses showed that autophagy-related GO terms or signaling pathways were enriched. GSEA analyses also revealed distinct differences in the autophagy-related signaling pathways between the high- and low-risk groups. Several cancer- and autophagy-related pathways were enriched in the high-risk group, whereas immunomodulatory pathways were enriched in the low-risk group. This suggested that increased immunity correlates with improved prognosis. These results were concordant with the current understanding that autophagy is a critical modulator of BLCA progression [12, 13].
There are several limitations in our study. Firstly, our findings need to be further validated in other independent cohorts to determine the robustness of the autophagy-related lncRNA prognostic signature. Secondly, our study was based on a single cohort of 409 patients from the publicly available TCGA database. Moreover, samples belonging to BCLA patients with high-grade tumor (n = 385) were significantly larger than those with low-grade tumors (n = 21), which may have skewed our results and hence need to be further analyzed with larger and more even number of samples in the high-risk and low-risk groups. Finally, further investigations involving biochemical experiments such as immunohistochemistry, quantitative real-time PCR, and flow cytometry, and clinical data analyses are required to further confirm our findings.
In conclusion, our study showed that the autophagy-related lncRNA prognostic signature accurately predicts the survival outcomes of BCLA patients with BLCA and distinguishes them into high- and low-risk groups. We also established and validated a prognostic nomogram by combining the autophagy-related lncRNA prognostic signature and other clinicopathological features. Our study demonstrated that this nomogram can provide an individualized and accurate survival prediction. Our study also suggests that these 5 autophagy-related lncRNAs are potential prognostic and diagnostic biomarkers as well as promising targets for BCLA therapy.
Materials and Methods
Patient data acquisition
The raw RNA-sequencing (RNA-seq) data and clinical information of 409 BLCA patients was downloaded from The Cancer Genome Atlas (TCGA) data portal (https://tcgadata.nci.nih.gov/tcga/). The Ensembl human genome browser, GRCh38.p13 (http://asia.ensembl.org/index.html) was used to annotate and classify the lncRNAs and protein-coding genes [36]. Patient samples were excluded (n = 16) if survival times of patients were less than 30 days to eliminate non-cancer related deaths. In addition, patients with incomplete clinical data (grade stage, n = 3; AJCC stage, n = 2) were excluded from the study. Since the data was obtained from a public database, approval from the Ethics committee or written informed consent from patients was not required.
Construction of the prognostic signature
The univariate Cox regression model was used to identify autophagy-related lncRNAs whose expression levels were significantly associated (P < 0.05) with the overall survival (OS) of the BLCA patient cohort. The hazard ratios (HRs) were used to identify risk-related lncRNAs (HR > 1) and protective lncRNAs (HR < 1). Subsequently, the candidate autophagy-related lncRNAs were subjected to multivariate Cox regression analysis to evaluate their contribution as independent prognostic factors in patient survival. Thus, we identified five target autophagy-related lncRNAs as candidates for the prognostic signature model, which was constructed based on a linear combination of the lncRNA expression levels and regression coefficients obtained from the multivariate Cox regression model. The optimal lncRNA prognostic signature was selected based on the lowest Akaike information criterion (AIC) value for further analysis. The computational formula used to determine the risk score for each patient based on this prognostic signature model was as follows:
Evaluation of the prognostic signature
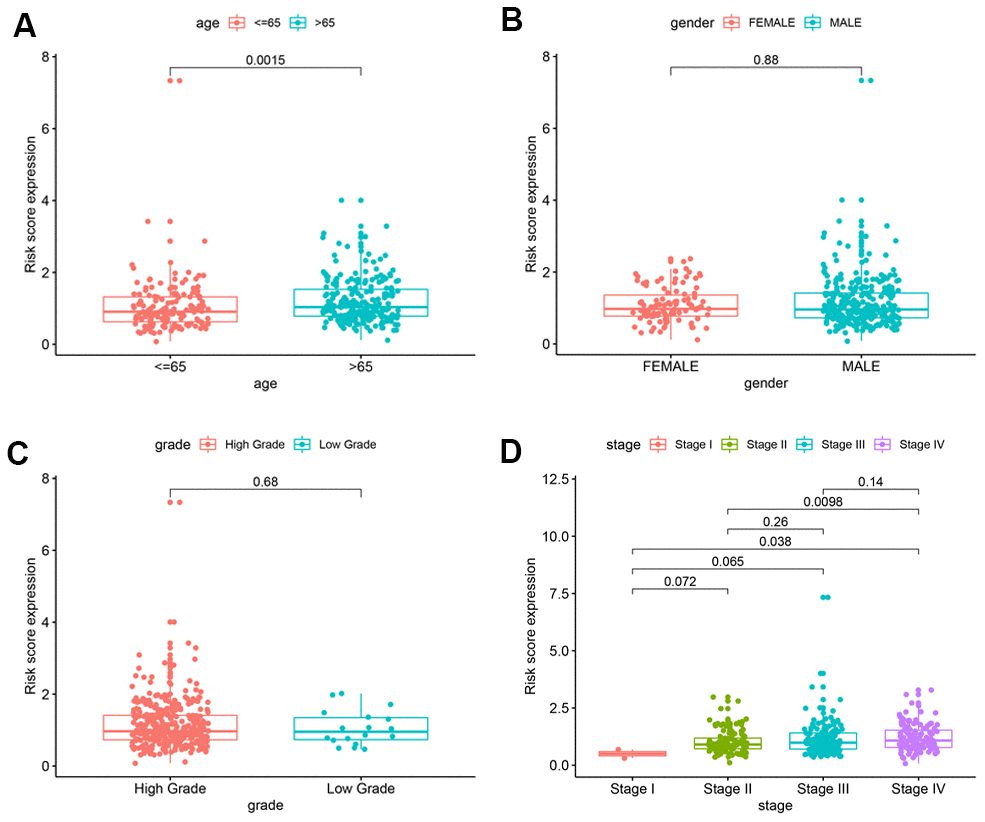
The BCLA patients were classified into high-risk or low-risk groups based on their prognostic risk score by using the median risk score as a cut-off point. The Kaplan–Meier survival curve and two-sided log-rank test was used to compare the overall survival (OS) of the high- and low-risk group patients. Principal component analysis (PCA) was performed to visualize gene expression patterns in the patient samples from the two groups. The receiver-operating characteristic (ROC) curves were applied to evaluate the diagnostic efficacy of each clinicopathological characteristic and the prognostic signature. Stratified survival analysis was performed to examine the accuracy of the prognostic signature in predicting patient survival outcomes. Furthermore, univariate and multivariate Cox regression analyses were performed to evaluate whether the risk score was independent of other clinical variables such as age, gender, grade, AJCC stage, T stage and N stage in determining the prognosis of the BCLA patients. M stage was not analyzed because the data was missing for several patients. P < 0.05 was considered statistically significant.
Establishment and validation of nomogram
We constructed a nomogram by integrating traditional clinical variables such as age, gender, grade, AJCC stage, T stage and N stage as well as the risk score derived from the prognostic signature to analyze the probable 1-, 3-, and 5-year OS of the BLCA patients. We then used the concordance index (C-index) to evaluate the discrimi-nation and predictive ability of the nomogram. The range of the C-index value was 0.5 to 1.0. A higher C-index indicates greater discrimination ability of the predicting model. Furthermore, calibration curves of the nomogram were generated to examine the concordance between pre-dicted survival and observed survival after bias correction.
Construction of the LncRNA-mRNA co-expression network
The mRNA-lncRNA co-expression network was constructed to analyze the correlation between the autophagy-related lncRNAs and their target mRNAs. Pearson correlation coefficients were calculated to identify the mRNAs that are significantly associated with their target lncRNAs based on the absolute threshold coefficient value > 0.3. The lncRNA-mRNA co-expression network was constructed and visualized using the Cytoscape software (version 3.7.2, http://www.cytoscape.org/).
Functional enrichment analysis
The lncRNA-related mRNAs were subjected to gene ontology (GO) enrichment analysis to identify the biological processes, molecular functions, and cellular components associated with the lncRNAs. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was used to determine the main signaling pathways regulated by these lncRNAs. P < 0.05 was considered statistically significant.
Gene set enrichment analysis (GSEA)
The genome wide expression profiles of the BCLA patients were subjected to gene set enrichment analysis (GSEA; http://www.broadinstitute.org/gsea) to determine the genes that are differentially expressed between the high- and low-risk group patients [38]. The gene sets were filtered using the maximum and minimum gene set size of 500 and 15 genes, respectively. The enriched gene sets were obtained based on a P value < 0.05 and a false discovery rate (FDR) value < 0.25 after performing 1,000 permutations.
Statistical analysis
The data was processed using the PERL programming language (Version 5.30.2, http://www.perl.org). All statistical analyses were performed using the R software (version 3.6.2, https://www.r-project.org/). P < 0.05 was regarded as statistically significant.
Author Contributions
SZL developed the study concept and design, performed data acquisition and analysis, and drafted the manuscript; JCY performed the bioinformatics and the statistical data analysis; XCT constructed the figures and tables; LTC was responsible for the integrity of the entire study and manuscript review. All authors read and approved the final manuscript.
Conflicts of Interest
The authors declare that there are no conflicts of interest.
References
- 1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017; 67:7–30. https://doi.org/10.3322/caac.21387 [PubMed]
- 2. Stenzl A, Cowan NC, De Santis M, Jakse G, Kuczyk MA, Merseburger AS, Ribal MJ, Sherif A, Witjes JA. The updated EAU guidelines on muscle-invasive and metastatic bladder cancer. Eur Urol. 2009; 55:815–25. https://doi.org/10.1016/j.eururo.2009.01.002 [PubMed]
- 3. Kamat AM, Hahn NM, Efstathiou JA, Lerner SP, Malmström PU, Choi W, Guo CC, Lotan Y, Kassouf W. Bladder cancer. Lancet. 2016; 388:2796–810. https://doi.org/10.1016/S0140-6736(16)30512-8 [PubMed]
- 4. Santoni M, Catanzariti F, Minardi D, Burattini L, Nabissi M, Muzzonigro G, Cascinu S, Santoni G. Pathogenic and diagnostic potential of BLCA-1 and BLCA-4 nuclear proteins in urothelial cell carcinoma of human bladder. Adv Urol. 2012; 2012:397412. https://doi.org/10.1155/2012/397412 [PubMed]
- 5. Mizushima N, Komatsu M. Autophagy: renovation of cells and tissues. Cell. 2011; 147:728–41. https://doi.org/10.1016/j.cell.2011.10.026 [PubMed]
- 6. Kimmelman AC, White E. Autophagy and tumor metabolism. Cell Metab. 2017; 25:1037–43. https://doi.org/10.1016/j.cmet.2017.04.004 [PubMed]
- 7. Menzies FM, Fleming A, Rubinsztein DC. Compromised autophagy and neurodegenerative diseases. Nat Rev Neurosci. 2015; 16:345–57. https://doi.org/10.1038/nrn3961 [PubMed]
- 8. Shirakabe A, Ikeda Y, Sciarretta S, Zablocki DK, Sadoshima J. Aging and autophagy in the heart. Circ Res. 2016; 118:1563–76. https://doi.org/10.1161/CIRCRESAHA.116.307474 [PubMed]
- 9. Matsuzawa-Ishimoto Y, Hwang S, Cadwell K. Autophagy and inflammation. Annu Rev Immunol. 2018; 36:73–101. https://doi.org/10.1146/annurev-immunol-042617-053253 [PubMed]
- 10. Singh SS, Vats S, Chia AY, Tan TZ, Deng S, Ong MS, Arfuso F, Yap CT, Goh BC, Sethi G, Huang RY, Shen HM, Manjithaya R, Kumar AP. Dual role of autophagy in hallmarks of cancer. Oncogene. 2018; 37:1142–58. https://doi.org/10.1038/s41388-017-0046-6 [PubMed]
- 11. Song S, Tan J, Miao Y, Li M, Zhang Q. Crosstalk of autophagy and apoptosis: involvement of the dual role of autophagy under ER stress. J Cell Physiol. 2017; 232:2977–84. https://doi.org/10.1002/jcp.25785 [PubMed]
- 12. Zhu J, Tian Z, Li Y, Hua X, Zhang D, Li J, Jin H, Xu J, Chen W, Niu B, Wu XR, Comincini S, Huang H, Huang C. ATG7 promotes bladder cancer invasion via autophagy-mediated increased ARHGDIB mRNA stability. Adv Sci (Weinh). 2019; 6:1801927. https://doi.org/10.1002/advs.201801927 [PubMed]
- 13. Zeng Q, Liu J, Cao P, Li J, Liu X, Fan X, Liu L, Cheng Y, Xiong W, Li J, Bo H, Zhu Y, Yang F, et al. Inhibition of REDD1 sensitizes bladder urothelial carcinoma to paclitaxel by inhibiting autophagy. Clin Cancer Res. 2018; 24:445–59. https://doi.org/10.1158/1078-0432.CCR-17-0419 [PubMed]
- 14. Alexander RP, Fang G, Rozowsky J, Snyder M, Gerstein MB. Annotating non-coding regions of the genome. Nat Rev Genet. 2010; 11:559–71. https://doi.org/10.1038/nrg2814 [PubMed]
- 15. Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012; 482:339–46. https://doi.org/10.1038/nature10887 [PubMed]
- 16. Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet. 2016; 17:47–62. https://doi.org/10.1038/nrg.2015.10 [PubMed]
- 17. Yan X, Hu Z, Feng Y, Hu X, Yuan J, Zhao SD, Zhang Y, Yang L, Shan W, He Q, Fan L, Kandalaft LE, Tanyi JL, et al. Comprehensive genomic characterization of long non-coding RNAs across human cancers. Cancer Cell. 2015; 28:529–40. https://doi.org/10.1016/j.ccell.2015.09.006 [PubMed]
- 18. Ying L, Huang Y, Chen H, Wang Y, Xia L, Chen Y, Liu Y, Qiu F. Downregulated MEG3 activates autophagy and increases cell proliferation in bladder cancer. Mol Biosyst. 2013; 9:407–11. https://doi.org/10.1039/c2mb25386k [PubMed]
- 19. Yuan P, Cao W, Zang Q, Li G, Guo X, Fan J. The HIF-2α-MALAT1-miR-216b axis regulates multi-drug resistance of hepatocellular carcinoma cells via modulating autophagy. Biochem Biophys Res Commun. 2016; 478:1067–73. https://doi.org/10.1016/j.bbrc.2016.08.065 [PubMed]
- 20. Huang F, Chen W, Peng J, Li Y, Zhuang Y, Zhu Z, Shao C, Yang W, Yao H, Zhang S. LncRNA PVT1 triggers cyto-protective autophagy and promotes pancreatic ductal adenocarcinoma development via the miR-20a-5p/ULK1 axis. Mol Cancer. 2018; 17:98. https://doi.org/10.1186/s12943-018-0845-6 [PubMed]
- 21. Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009; 37:1–13. https://doi.org/10.1093/nar/gkn923 [PubMed]
- 22. Desany B, Zhang Z. Bioinformatics and cancer target discovery. Drug Discov Today. 2004; 9:795–802. https://doi.org/10.1016/S1359-6446(04)03224-6 [PubMed]
- 23. Balachandran VP, Gonen M, Smith JJ, DeMatteo RP. Nomograms in oncology: more than meets the eye. Lancet Oncol. 2015; 16:e173–80. https://doi.org/10.1016/S1470-2045(14)71116-7 [PubMed]
- 24. Wang KJ, Wang C, Dai LH, Yang J, Huang H, Ma XJ, Zhou Z, Yang ZY, Xu WD, Hua MM, Lu X, Zeng SX, Wang HQ, et al. Targeting an autocrine regulatory loop in cancer stem-like cells impairs the progression and chemotherapy resistance of bladder cancer. Clin Cancer Res. 2019; 25:1070–86. https://doi.org/10.1158/1078-0432.CCR-18-0586 [PubMed]
- 25. Li F, Guo H, Yang Y, Feng M, Liu B, Ren X, Zhou H. Autophagy modulation in bladder cancer development and treatment (review). Oncol Rep. 2019; 42:1647–55. https://doi.org/10.3892/or.2019.7286 [PubMed]
- 26. Wang SS, Chen G, Li SH, Pang JS, Cai KT, Yan HB, Huang ZG, He RQ. Identification and validation of an individualized autophagy-clinical prognostic index in bladder cancer patients. Onco Targets Ther. 2019; 12:3695–712. https://doi.org/10.2147/OTT.S197676 [PubMed]
- 27. Martens-Uzunova ES, Böttcher R, Croce CM, Jenster G, Visakorpi T, Calin GA. Long noncoding RNA in prostate, bladder, and kidney cancer. Eur Urol. 2014; 65:1140–51. https://doi.org/10.1016/j.eururo.2013.12.003 [PubMed]
- 28. Bendifallah S, Daraï E, Ballester M. Predictive modeling: a new paradigm for managing endometrial cancer. Ann Surg Oncol. 2016; 23:975–88. https://doi.org/10.1245/s10434-015-4924-2 [PubMed]
- 29. Yorimitsu T, Klionsky DJ. Autophagy: molecular machinery for self-eating. Cell Death Differ. 2005 (Suppl 2); 12:1542–52. https://doi.org/10.1038/sj.cdd.4401765 [PubMed]
- 30. Kreuzaler P, Watson CJ. Killing a cancer: what are the alternatives? Nat Rev Cancer. 2012; 12:411–24. https://doi.org/10.1038/nrc3264 [PubMed]
- 31. White E, DiPaola RS. The double-edged sword of autophagy modulation in cancer. Clin Cancer Res. 2009; 15:5308–16. https://doi.org/10.1158/1078-0432.CCR-07-5023 [PubMed]
- 32. Mathew R, Karantza-Wadsworth V, White E. Role of autophagy in cancer. Nat Rev Cancer. 2007; 7:961–67. https://doi.org/10.1038/nrc2254 [PubMed]
- 33. Xu Z, Han X, Ou D, Liu T, Li Z, Jiang G, Liu J, Zhang J. Targeting PI3K/AKT/mTOR-mediated autophagy for tumor therapy. Appl Microbiol Biotechnol. 2020; 104:575–87. https://doi.org/10.1007/s00253-019-10257-8 [PubMed]
- 34. Lee MJ, Lee JH, Rubinsztein DC. Tau degradation: the ubiquitin-proteasome system versus the autophagy-lysosome system. Prog Neurobiol. 2013; 105:49–59. https://doi.org/10.1016/j.pneurobio.2013.03.001 [PubMed]
- 35. Wang CZ, Yan GX, Dong DS, Xin H, Liu ZY. LncRNA-ATB promotes autophagy by activating yes-associated protein and inducing autophagy-related protein 5 expression in hepatocellular carcinoma. World J Gastroenterol. 2019; 25:5310–22. https://doi.org/10.3748/wjg.v25.i35.5310 [PubMed]
- 36. Cunningham F, Achuthan P, Akanni W, Allen J, Amode MR, Armean IM, Bennett R, Bhai J, Billis K, Boddu S, Cummins C, Davidson C, Dodiya KJ, et al. Ensembl 2019. Nucleic Acids Res. 2019; 47:D745–51. https://doi.org/10.1093/nar/gky1113 [PubMed]
- 37. Moussay E, Kaoma T, Baginska J, Muller A, Van Moer K, Nicot N, Nazarov PV, Vallar L, Chouaib S, Berchem G, Janji B. The acquisition of resistance to TNFα in breast cancer cells is associated with constitutive activation of autophagy as revealed by a transcriptome analysis using a custom microarray. Autophagy. 2011; 7:760–70. https://doi.org/10.4161/auto.7.7.15454 [PubMed]
- 38. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005; 102:15545–50. https://doi.org/10.1073/pnas.0506580102 [PubMed]