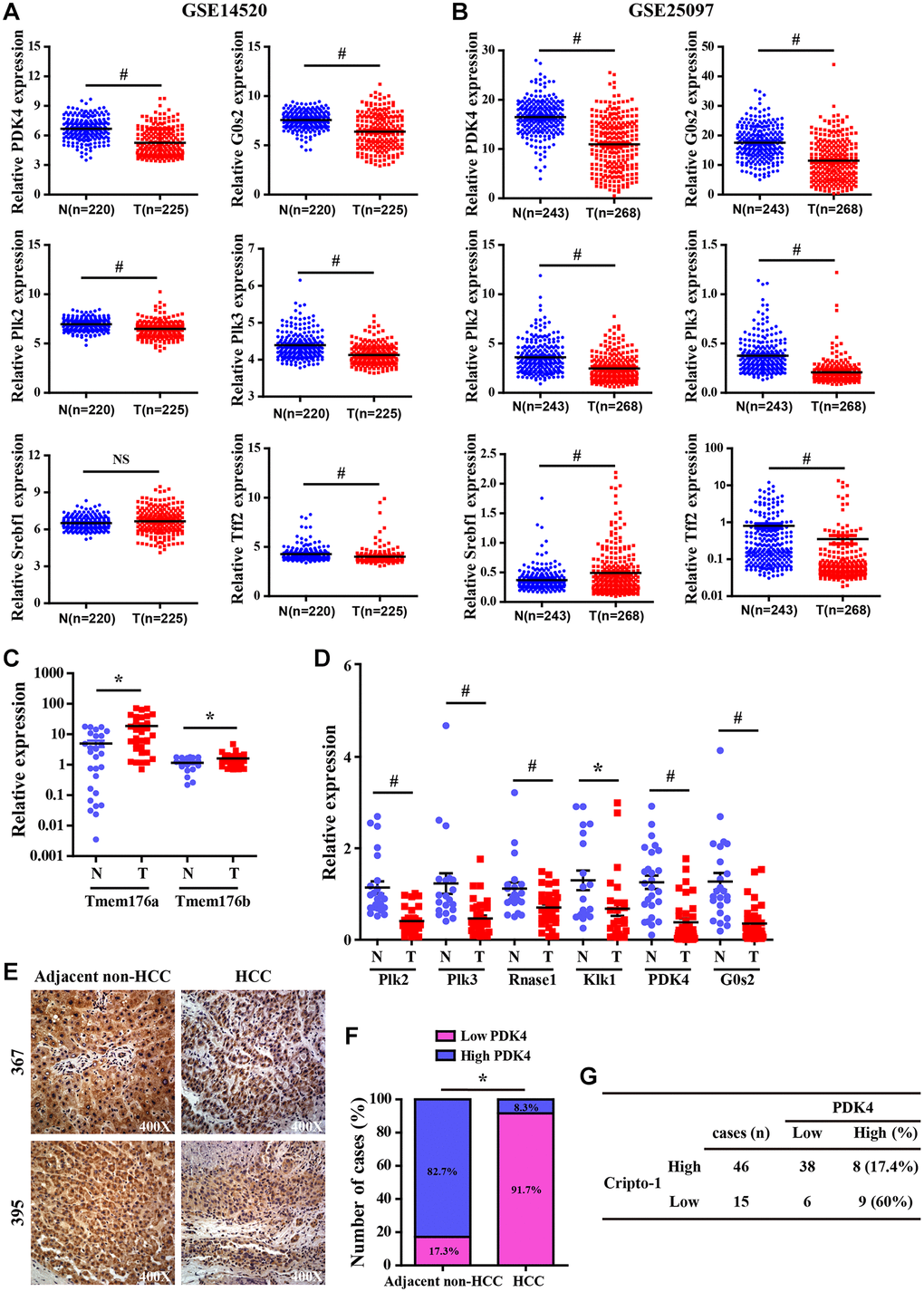
Figure 7.Validation analysis of differentially expressed genes from the liver of RCLG/Alb-Cre mice in HCC tissues. (A–B) The expression levels of CR-1 in HCC (T) and adjacent non-cancerous (N) liver tissue biopsies derived from NCBI-GEO datasets (GSE14520 and GSE25097). (C–D) qRT-PCR analysis shows expression of the indicated genes (selected from Figure 6A–6D) in primary HCC (T) and matched non-tumor liver (N) tissues. (E) Representative IHC images show PDK4 expression in HCC and adjacent non-tumor liver tissue biopsies. (F) IHC assay results show the percentage of HCC and adjacent non-tumor liver tissue biopsies with high or low PDK4 expression. As shown, PDK4 expression was significantly lower in the HCC tissue biopsies than in the adjacent non-tumor liver tissues. (G) The correlation analysis between CR-1 and PDK4 expression in HCC specimens based on PDK4 IHC data.