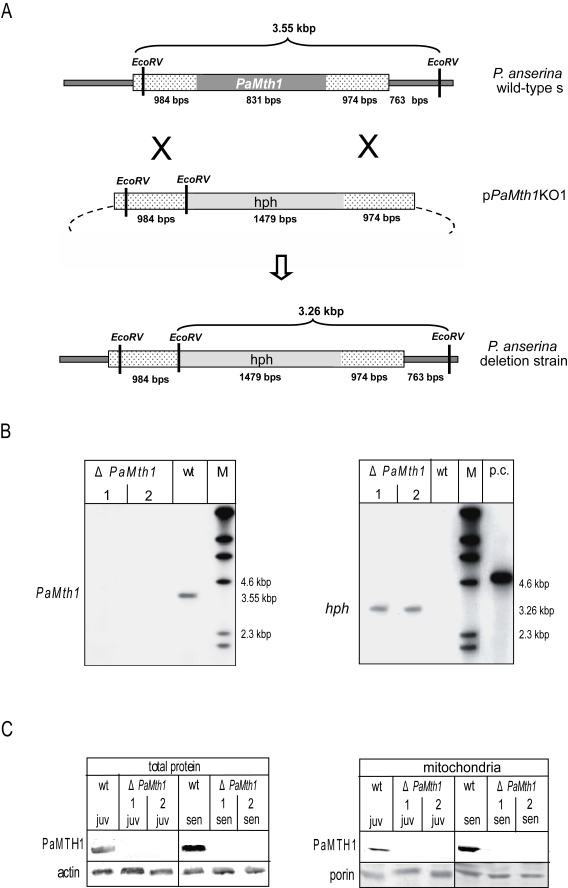
Figure 1.Construction and
validation of a PaMth1 deletion strain of P. anserina. (A)
Physical maps and sizes of the restriction products for the genomic region
bearing PaMth1 and the recombined version with the hygromycin B
resistance cassette. Reading frames of the two genes are indicated in grey.
The genomic sequences flanking PaMth1 are indicated by punctuation.
Restriction sites of EcoRV are indicated. (B) Southern blot
analysis of Eco RV digested wild-type strain s genomic DNA and
genomic DNA of two secondary transformants isolated from a primary deletion
strain. The PaMth1 gene-specific probe (left panel) detects the 3.55
kbp fragment only in the sample of the wild-type s (wt) but not in the
samples of the deletions strains (ΔPaMth1). The hygromycin B
resistance gene-specific probe (right panel) detects the 3.26 kbp fragment
only in the sample of the deletions strains. (C) Western blot
analysis verifying the successful construction of a PaMth1 deletion
strain using total- and mitochondrial protein samples of juvenile and
senescent P. anserina wild-type strain s and the secondary
transformants of the deletion strain, respectively. The PaMTH1 specific
antibody detects PaMTH1 in the samples of the wild-type s strains but not
in samples of the deletion strains. As loading controls an actin specific
antibody for total proteins and a porin specific antibody for the
mitochondrial proteins were used.