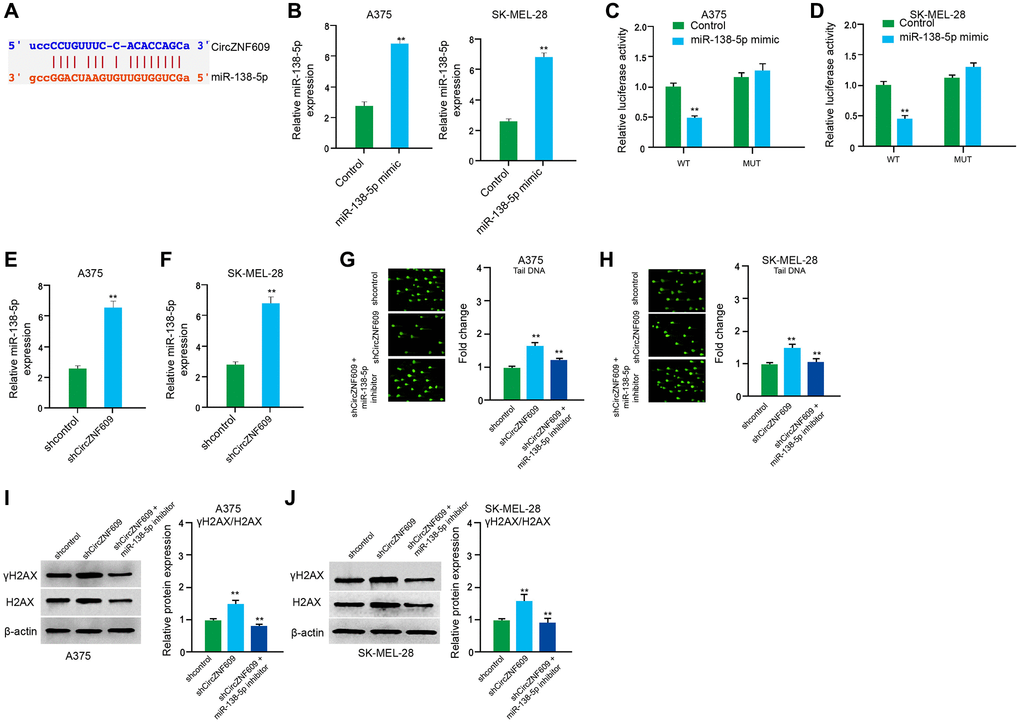
Figure 4.CircZNF609 inhibits DNA damage by sponging miR-138-5p in melanoma cells. (A) The interaction of circZNF609 and miR-138-5p was analyzed by bioinformatic analysis based on ENCORI (starbase.sysu.edu.cn). (B–D) The A375 and SK-MEL-28 cells treated with control mimic or miR-138-5p mimics. The expression levels of miR-138-5p were tested by qPCR in the cells. (C and D) Luciferase activities of circZNF609 (circZNF609 WT) and circZNF609 with the miR-138-5p-binding site mutant (circZNF609 MUT) were examined by luciferase reporter gene assays in the cells. (E and F) The A375 and SK-MEL-28 cells were treated with the circZNF609 shRNA or control shRNA. The expression of miR-138-5p was analyzed by qPCR assays in the cells. (G–J) The A375 and SK-MEL-28 cells were treated with the control shRNA or circZNF609 shRNA, or co-treated with circZNF609 shRNA and miR-138-5p inhibitor. (G and H) The DNA damage was analyzed by comet assays in the cells. (I and J) The protein expression of H2AX, γH2AX and β-actin was determined by Western blot analysis in the cells. The results of Western blot analysis were quantified by ImageJ software. Data are presented as mean ± SD. Statistic significant differences were indicated: *P < 0.05, **P < 0.01.