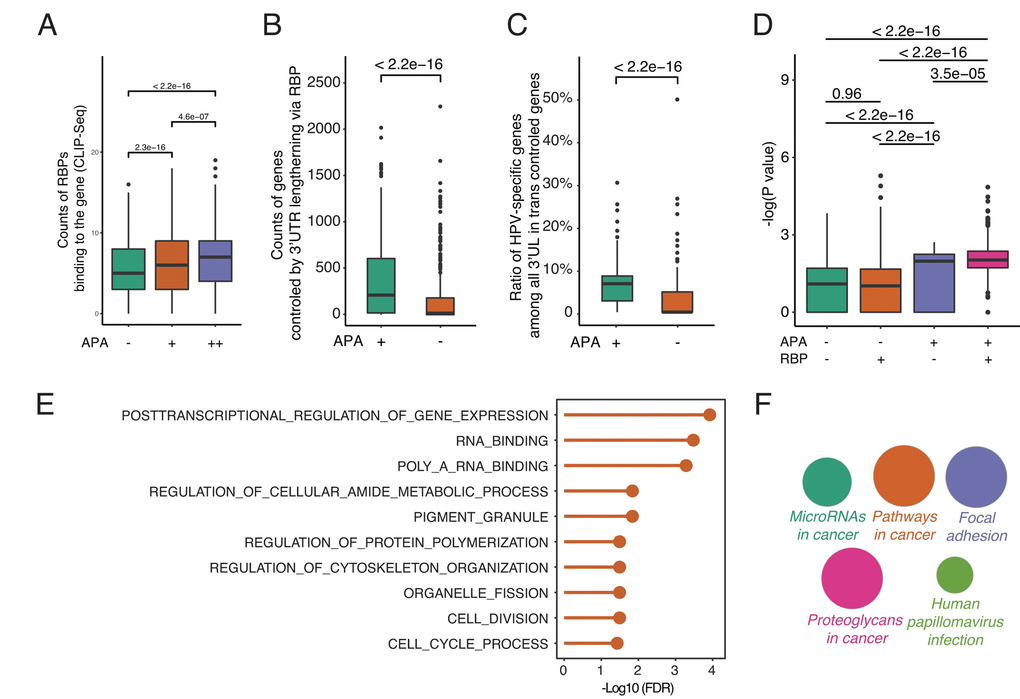
Figure 3.By releasing intermediate RNA-binding proteins (RBPs), 3’UL genes indirectly impact paired genes in trans. (A) Differential-APA genes are easier to be attacked by RBPs (Wilcoxon test, data from CLIP-seq). APA+ group and APA++ group represent different differential APA level (P < 0.05 and P < 0.001, respectively). (B) 3’UL controls more paired genes in trans compared with non-3’UL APA events. This computation was generated by randomly selecting 1,000 3’UL-gene pairs. In order to retrieve high-confidence interactions, we limit the number of intermediate RBPs > 3. (C) 3’UL-in-trans-paired genes show significantly higher ratio of HPV-specific. This computation was generated by randomly selecting 1,000 3’UL-gene pairs with the number of intermediate RBPs > 3. (D) Through intermediate RBPs, 3’UL controls paired genes in trans. APA+RBP+ group shows the most significant HPV-specific trend. The bigger the Y-axis value is, the group harbors more HPV-specific trend. The Y-axis marks the -log (P-value) of differential expression between HPV+ and HPV- samples through EdgeR. The P-value between 4 groups was computed through the Wilcoxon test. This computation was generated by randomly selecting 1,000 3’UL-gene pairs. RBP+/RBP- was defined as higher/lower RBP-binding confidence level (intermediate RBPs > 3 and ≤ 3, respectively). In the RBP+ group, 3’UL impacts in-trans-paired genes with higher confidence. (E) Biological process enrichment analysis of HUB-3’UL in-trans-paired genes (hypergeometric test, retrieved from Molecular Signature Database). The cell cycle is the most top ranking for enrichment term. HUB-3’UL was defined as the 3’UL genes with interacting RBPs ≥ 15. (F) Pathway enrichment of HUB-3’UL in-trans-paired genes. CLUEGO package was used based on KEGG database. The size of each bubble represents the enrichment level of each term.