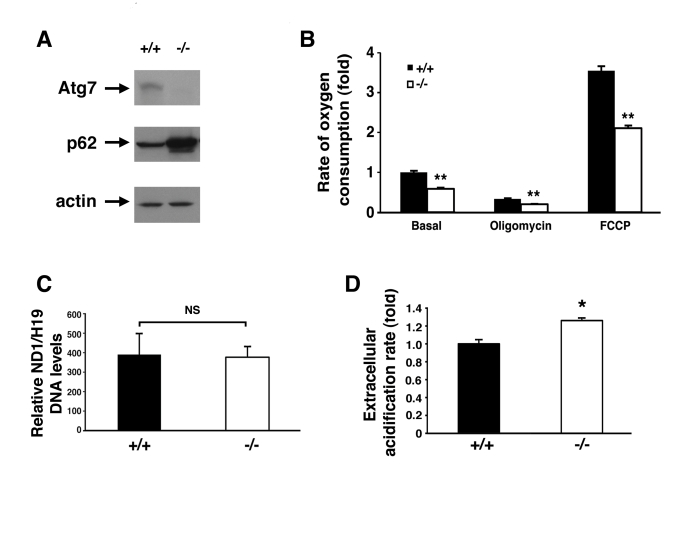
Figure 2.Alterations in the energetics of Atg7 -/- MEFs.
(A) Western blot
analysis of wild type (+/+) or Atg7-/- MEFs for the expression
of Atg7, p62 and actin (loading control). (B) Measurement of oxygen
consumption for WT and Atg7-/- MEFs under basal conditions,
following the addition of the mitochondrial electron chain inhibitor
oligomycin (0.5 μM), or in the presence of the mitochondrial uncoupler FCCP
(1 μM), to determine maximal oxidative capacity. Shown is the average fold
change +/- SEM in oxygen consumption (WT MEFs basal respiration=1) obtained
from 5 experiments each performed in triplicate. (C) Assessment of
mitochondrial number in WT or Atg7-/- MEFs. DNA was isolated
from WT (n=3 independent WT MEF cell isolates) and Atg7-/- MEFs (n=3
independent Atg7-/- MEF cell isolates) and quantitative PCR
analysis performed for the mitochondrial-encoded gene ND1 and the
nuclear-encoded gene H19. (D) Relative extracellular acidification
rates indicating lactic acid production and hence glyolytic rates in WT or
Atg7-/- MEFs. Shown is the average +/- SEM fold change in
lactic acid production from 8 experiments each performed in triplicate. *
p≤0.05; ** p≤0.01.